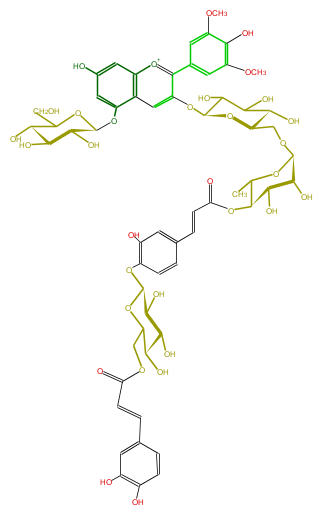
Mol:FL7AAIGL0026
From Metabolomics.JP
Copyright: ARM project http://www.metabolome.jp/
91 99 0 0 0 0 0 0 0 0999 V2000
0.7920 5.6750 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.6171 5.6750 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0296 6.3895 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.6169 7.1038 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7921 7.1039 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.3797 6.3894 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0295 4.9605 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.8544 4.9605 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.2669 5.6751 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.8544 6.3894 0.0000 O 0 3 0 0 0 0 0 0 0 0 0 0
4.0528 5.6750 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.4472 4.9919 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.2358 4.9919 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.6301 5.6750 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.2358 6.3580 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.4471 6.3580 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
6.4402 5.6750 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.4547 7.6883 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.4560 5.2742 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.2405 4.2919 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.3878 6.6624 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.1106 5.7375 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1453 5.7603 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.3031 5.2881 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5801 6.2131 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.5456 6.1904 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.2173 6.5805 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.8178 6.1207 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.7460 4.9191 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.4691 3.9492 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.5323 3.7156 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3681 3.2320 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.8410 2.3902 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.7780 2.6238 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.9421 3.1074 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.9492 4.2616 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.8356 3.1237 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
6.2779 1.9056 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.6110 2.0485 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.5932 1.3481 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.8374 0.6832 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.3546 -0.1529 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.2831 0.1124 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.2171 -0.1324 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.7001 0.7037 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.7716 0.4384 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.2644 -0.8351 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.8571 -0.6378 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.0609 0.5419 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.5960 0.1024 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.0796 0.7541 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.4889 1.4629 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.3399 0.7540 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.9421 0.0652 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.2178 0.0652 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.1807 0.7554 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.9777 0.7554 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.3762 0.0652 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.9777 -0.6251 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.1806 -0.6251 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1358 1.1895 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.8658 0.0652 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.9358 -3.0090 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.0383 -2.6529 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.1446 -1.6932 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.7470 -0.8133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.6447 -1.1692 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.5383 -2.1291 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.8108 -3.8805 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.2406 -3.3988 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.4461 -1.2421 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.1566 -2.5090 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.3420 -3.4685 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.2182 -3.4953 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.7461 -2.6921 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.8558 -4.2150 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.3040 -5.0101 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.8266 -5.8101 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.6474 -5.8147 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-6.0537 -6.5278 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.6392 -7.2363 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.8186 -7.2316 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.4122 -6.5185 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-6.5929 -6.5278 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-6.0915 -7.6883 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.8068 4.3298 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
6.5929 4.3300 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.0291 7.1615 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.9124 7.6539 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.6246 7.0314 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
6.4282 7.0315 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
2 7 1 0 0 0 0
7 8 2 0 0 0 0
8 9 1 0 0 0 0
9 10 2 0 0 0 0
10 3 1 0 0 0 0
9 11 1 0 0 0 0
11 12 2 0 0 0 0
12 13 1 0 0 0 0
13 14 2 0 0 0 0
14 15 1 0 0 0 0
15 16 2 0 0 0 0
16 11 1 0 0 0 0
14 17 1 0 0 0 0
5 18 1 0 0 0 0
1 19 1 0 0 0 0
8 20 1 0 0 0 0
21 22 1 1 0 0 0
22 23 1 1 0 0 0
24 23 1 1 0 0 0
24 25 1 0 0 0 0
25 26 1 0 0 0 0
26 21 1 0 0 0 0
21 27 1 0 0 0 0
22 28 1 0 0 0 0
23 29 1 0 0 0 0
24 19 1 0 0 0 0
30 31 1 0 0 0 0
31 32 1 1 0 0 0
33 32 1 1 0 0 0
34 33 1 1 0 0 0
34 35 1 0 0 0 0
35 30 1 0 0 0 0
30 36 1 0 0 0 0
35 37 1 0 0 0 0
34 38 1 0 0 0 0
33 39 1 0 0 0 0
39 40 1 0 0 0 0
31 20 1 0 0 0 0
41 42 1 0 0 0 0
42 43 1 1 0 0 0
44 43 1 1 0 0 0
45 44 1 1 0 0 0
45 46 1 0 0 0 0
46 41 1 0 0 0 0
40 45 1 0 0 0 0
44 47 1 0 0 0 0
43 48 1 0 0 0 0
41 49 1 0 0 0 0
42 50 1 0 0 0 0
50 51 1 0 0 0 0
51 52 2 0 0 0 0
51 53 1 0 0 0 0
53 54 2 0 0 0 0
54 55 1 0 0 0 0
55 56 2 0 0 0 0
56 57 1 0 0 0 0
57 58 2 0 0 0 0
58 59 1 0 0 0 0
59 60 2 0 0 0 0
60 55 1 0 0 0 0
57 61 1 0 0 0 0
81 85 1 0 0 0 0
58 62 1 0 0 0 0
63 64 1 1 0 0 0
64 65 1 1 0 0 0
66 65 1 1 0 0 0
66 67 1 0 0 0 0
67 68 1 0 0 0 0
68 63 1 0 0 0 0
63 69 1 0 0 0 0
64 70 1 0 0 0 0
65 71 1 0 0 0 0
66 62 1 0 0 0 0
68 72 1 0 0 0 0
72 73 1 0 0 0 0
73 74 1 0 0 0 0
74 75 2 0 0 0 0
74 76 1 0 0 0 0
76 77 2 0 0 0 0
77 78 1 0 0 0 0
78 79 2 0 0 0 0
79 80 1 0 0 0 0
80 81 2 0 0 0 0
81 82 1 0 0 0 0
82 83 2 0 0 0 0
83 78 1 0 0 0 0
80 84 1 0 0 0 0
86 87 1 0 0 0 0
13 86 1 0 0 0 0
88 89 1 0 0 0 0
26 88 1 0 0 0 0
90 91 1 0 0 0 0
15 90 1 0 0 0 0
M CHG 1 10 1
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 86 87
M SBL 1 1 95
M SMT 1 OCH3
M SBV 1 95 -0.5710 0.6621
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 88 89
M SBL 2 1 97
M SMT 2 ^ CH2OH
M SBV 2 97 0.4835 -0.9711
M STY 1 3 SUP
M SLB 1 3 3
M SAL 3 2 90 91
M SBL 3 1 99
M SMT 3 OCH3
M SBV 3 99 -0.3888 -0.6734
S SKP 5
ID FL7AAIGL0026
FORMULA C59H67O32
EXACTMASS 1287.3615450480002
AVERAGEMASS 1288.14408
SMILES OC(C1OC(=O)C=Cc(c9)cc(O)c(c9)OC(O7)C(C(O)C(C7COC(=O)C=Cc(c8)ccc(c(O)8)O)O)O)C(C(OCC(C6O)OC(C(C6O)O)Oc(c(c(c5)cc(c(O)c5OC)OC)4)cc(c([o+1]4)3)c(cc(O)c3)OC(O2)C(O)C(O)C(C(CO)2)O)OC1C)O
M END