Mol:FL5FACGL0067
From Metabolomics.JP
Copyright: ARM project http://www.metabolome.jp/
67 73 0 0 0 0 0 0 0 0999 V2000
-4.0675 1.4336 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.0675 0.7912 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.5112 0.4700 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.9549 0.7912 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.9549 1.4336 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.5112 1.7547 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.3986 0.4700 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.8423 0.7912 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.8423 1.4336 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.3986 1.7547 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.3986 -0.0308 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.2862 1.7546 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.7193 1.4273 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.1523 1.7546 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.1523 2.4093 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.7193 2.7366 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.2862 2.4093 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.6236 1.7546 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.4145 2.7366 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.4482 0.4023 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.5112 -0.1721 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.7193 3.3911 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.1900 -1.5954 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0
-1.6079 -1.4451 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
-1.6357 -0.8504 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
-1.3450 -0.3372 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
-1.9835 -0.5626 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.9271 -1.1822 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
-2.1900 -2.0682 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.6079 -2.2015 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.1023 -0.8432 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.0024 -1.4641 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
-0.6412 -1.4641 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
-0.1821 -1.9083 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.0251 -2.5313 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
0.6138 -2.5313 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
0.1546 -2.0871 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
0.6131 -1.8733 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.9791 -2.7016 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.6045 0.5546 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.8508 0.9602 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.4217 0.9413 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.6154 1.4649 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.8653 1.8895 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.5987 2.4612 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.9301 2.9345 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.6859 3.4581 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.1103 3.5085 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.7790 3.0352 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0232 2.5116 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8663 4.0319 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.2710 -0.3342 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0
0.6727 -0.8644 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
1.2512 -0.6395 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
1.7681 -0.7159 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
1.4037 -0.2277 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
0.8977 -0.4948 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.4927 -0.7751 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.0465 -0.8949 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.5826 -1.1959 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.1263 -0.9228 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.9248 0.1642 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.0171 3.9312 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.5348 -0.7694 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.4350 -1.2048 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.1495 -3.0248 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.8640 -3.4373 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
1 18 1 0 0 0 0
15 19 1 0 0 0 0
8 20 1 0 0 0 0
3 21 1 0 0 0 0
16 22 1 0 0 0 0
23 24 1 1 0 0 0
24 25 1 1 0 0 0
26 25 1 1 0 0 0
26 27 1 0 0 0 0
27 28 1 0 0 0 0
28 23 1 0 0 0 0
23 29 1 0 0 0 0
24 30 1 0 0 0 0
25 31 1 0 0 0 0
26 20 1 0 0 0 0
32 33 1 0 0 0 0
33 34 1 1 0 0 0
34 35 1 1 0 0 0
36 35 1 1 0 0 0
36 37 1 0 0 0 0
37 32 1 0 0 0 0
37 38 1 0 0 0 0
36 39 1 0 0 0 0
33 31 1 0 0 0 0
40 41 1 0 0 0 0
41 42 2 0 0 0 0
41 43 1 0 0 0 0
43 44 2 0 0 0 0
44 45 1 0 0 0 0
45 46 2 0 0 0 0
46 47 1 0 0 0 0
47 48 2 0 0 0 0
48 49 1 0 0 0 0
49 50 2 0 0 0 0
50 45 1 0 0 0 0
48 51 1 0 0 0 0
52 53 1 1 0 0 0
53 54 1 1 0 0 0
55 54 1 1 0 0 0
55 56 1 0 0 0 0
56 57 1 0 0 0 0
57 52 1 0 0 0 0
52 58 1 0 0 0 0
53 59 1 0 0 0 0
54 60 1 0 0 0 0
58 32 1 0 0 0 0
55 61 1 0 0 0 0
56 62 1 0 0 0 0
62 40 1 0 0 0 0
47 63 1 0 0 0 0
28 64 1 0 0 0 0
64 65 1 0 0 0 0
35 66 1 0 0 0 0
66 67 1 0 0 0 0
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 66 67
M SBL 2 1 72
M SMT 2 CH2OH
M SVB 2 72 0.1495 -3.0248
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 64 65
M SBL 1 1 70
M SMT 1 CH2OH
M SVB 1 70 -2.5348 -0.7694
S SKP 8
ID FL5FACGL0067
KNApSAcK_ID C00005986
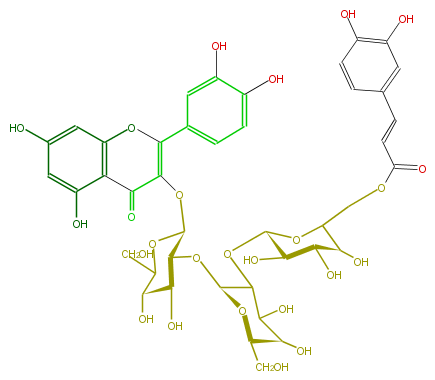
NAME Quercetin 3-(6''''-caffeylsophorotrioside)
CAS_RN 167638-64-4
FORMULA C42H46O25
EXACTMASS 950.2328170220001
AVERAGEMASS 950.79964
SMILES [C@H]([C@@H](O)4)(O)[C@@H]([C@H](OC(C(=O)5)=C(c(c7)cc(c(c7)O)O)Oc(c6)c5c(O)cc(O)6)OC4CO)O[C@@H](C1O[C@@H](O2)[C@@H]([C@@H]([C@@H](C(COC(=O)C=Cc(c3)cc(O)c(c3)O)2)O)O)O)O[C@@H]([C@@H](C1O)O)CO
M END