Mol:FL7AAIGL0018
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 52 56 0 0 0 0 0 0 0 0999 V2000 | + | 52 56 0 0 0 0 0 0 0 0999 V2000 |
| − | -1.3367 0.4603 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.3367 0.4603 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.3367 -0.1821 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.3367 -0.1821 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.7804 -0.5032 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7804 -0.5032 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.2241 -0.1821 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.2241 -0.1821 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.2241 0.4603 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.2241 0.4603 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.7804 0.7815 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7804 0.7815 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.3322 -0.5032 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.3322 -0.5032 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8885 -0.1821 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8885 -0.1821 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8885 0.4603 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8885 0.4603 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.3322 0.7815 0.0000 O 0 3 0 0 0 0 0 0 0 0 0 0 | + | 0.3322 0.7815 0.0000 O 0 3 0 0 0 0 0 0 0 0 0 0 |
| − | 1.4446 0.7814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.4446 0.7814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0116 0.4540 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0116 0.4540 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.5786 0.7814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5786 0.7814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.5786 1.4361 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5786 1.4361 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0116 1.7634 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0116 1.7634 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.4446 1.4361 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.4446 1.4361 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.8928 0.7814 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.8928 0.7814 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.4446 1.9361 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.4446 1.9361 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.7804 -1.1454 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7804 -1.1454 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3046 -0.9028 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3046 -0.9028 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.8075 -0.9265 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 2.8075 -0.9265 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 2.5393 -1.3911 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.5393 -1.3911 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 3.0551 -1.2437 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0551 -1.2437 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.5740 -1.3911 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 3.5740 -1.3911 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 3.8423 -0.9265 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 3.8423 -0.9265 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 3.3264 -1.0739 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 3.3264 -1.0739 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 2.4429 -0.5618 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.4429 -0.5618 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.4681 -0.5451 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.4681 -0.5451 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.2494 -0.9265 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.2494 -0.9265 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.6504 -1.5433 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 | + | -2.6504 -1.5433 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 |
| − | -2.2792 -2.0333 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | -2.2792 -2.0333 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | -1.7447 -1.8254 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | -1.7447 -1.8254 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | -1.2289 -1.8198 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | -1.2289 -1.8198 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | -1.6037 -1.4450 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6037 -1.4450 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.0713 -1.6918 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | -2.0713 -1.6918 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | -3.2413 -1.4916 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.2413 -1.4916 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.6296 -2.4945 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.6296 -2.4945 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4384 -2.3395 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4384 -2.3395 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.5082 -1.1897 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.5082 -1.1897 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.4206 -0.4639 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.4206 -0.4639 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.0763 -0.0853 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.0763 -0.0853 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.6433 -0.4126 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.6433 -0.4126 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.2101 -0.0854 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.2101 -0.0854 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.7770 -0.4126 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.7770 -0.4126 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.0763 0.4818 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.0763 0.4818 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.2101 0.5691 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.2101 0.5691 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.7088 -1.7870 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.7088 -1.7870 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.5655 -2.5994 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.5655 -2.5994 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2950 2.3395 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2950 2.3395 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.7364 3.2368 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.7364 3.2368 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.5786 0.7814 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.5786 0.7814 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.5786 0.7814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.5786 0.7814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 2 0 0 0 0 | + | 7 8 2 0 0 0 0 |
| − | 8 9 1 0 0 0 0 | + | 8 9 1 0 0 0 0 |
| − | 9 10 2 0 0 0 0 | + | 9 10 2 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 9 11 1 0 0 0 0 | + | 9 11 1 0 0 0 0 |
| − | 11 12 2 0 0 0 0 | + | 11 12 2 0 0 0 0 |
| − | 12 13 1 0 0 0 0 | + | 12 13 1 0 0 0 0 |
| − | 13 14 2 0 0 0 0 | + | 13 14 2 0 0 0 0 |
| − | 14 15 1 0 0 0 0 | + | 14 15 1 0 0 0 0 |
| − | 15 16 2 0 0 0 0 | + | 15 16 2 0 0 0 0 |
| − | 16 11 1 0 0 0 0 | + | 16 11 1 0 0 0 0 |
| − | 1 17 1 0 0 0 0 | + | 1 17 1 0 0 0 0 |
| − | 14 18 1 0 0 0 0 | + | 14 18 1 0 0 0 0 |
| − | 3 19 1 0 0 0 0 | + | 3 19 1 0 0 0 0 |
| − | 20 8 1 0 0 0 0 | + | 20 8 1 0 0 0 0 |
| − | 21 22 1 0 0 0 0 | + | 21 22 1 0 0 0 0 |
| − | 22 23 1 1 0 0 0 | + | 22 23 1 1 0 0 0 |
| − | 23 24 1 1 0 0 0 | + | 23 24 1 1 0 0 0 |
| − | 25 24 1 1 0 0 0 | + | 25 24 1 1 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 21 1 0 0 0 0 | + | 26 21 1 0 0 0 0 |
| − | 21 27 1 0 0 0 0 | + | 21 27 1 0 0 0 0 |
| − | 26 28 1 0 0 0 0 | + | 26 28 1 0 0 0 0 |
| − | 25 29 1 0 0 0 0 | + | 25 29 1 0 0 0 0 |
| − | 22 20 1 0 0 0 0 | + | 22 20 1 0 0 0 0 |
| − | 30 31 1 1 0 0 0 | + | 30 31 1 1 0 0 0 |
| − | 31 32 1 1 0 0 0 | + | 31 32 1 1 0 0 0 |
| − | 33 32 1 1 0 0 0 | + | 33 32 1 1 0 0 0 |
| − | 33 34 1 0 0 0 0 | + | 33 34 1 0 0 0 0 |
| − | 34 35 1 0 0 0 0 | + | 34 35 1 0 0 0 0 |
| − | 35 30 1 0 0 0 0 | + | 35 30 1 0 0 0 0 |
| − | 30 36 1 0 0 0 0 | + | 30 36 1 0 0 0 0 |
| − | 31 37 1 0 0 0 0 | + | 31 37 1 0 0 0 0 |
| − | 32 38 1 0 0 0 0 | + | 32 38 1 0 0 0 0 |
| − | 33 19 1 0 0 0 0 | + | 33 19 1 0 0 0 0 |
| − | 35 39 1 0 0 0 0 | + | 35 39 1 0 0 0 0 |
| − | 39 40 1 0 0 0 0 | + | 39 40 1 0 0 0 0 |
| − | 40 41 1 0 0 0 0 | + | 40 41 1 0 0 0 0 |
| − | 41 42 1 0 0 0 0 | + | 41 42 1 0 0 0 0 |
| − | 42 43 1 0 0 0 0 | + | 42 43 1 0 0 0 0 |
| − | 43 44 2 0 0 0 0 | + | 43 44 2 0 0 0 0 |
| − | 41 45 2 0 0 0 0 | + | 41 45 2 0 0 0 0 |
| − | 43 46 1 0 0 0 0 | + | 43 46 1 0 0 0 0 |
| − | 24 47 1 0 0 0 0 | + | 24 47 1 0 0 0 0 |
| − | 47 48 1 0 0 0 0 | + | 47 48 1 0 0 0 0 |
| − | 15 49 1 0 0 0 0 | + | 15 49 1 0 0 0 0 |
| − | 49 50 1 0 0 0 0 | + | 49 50 1 0 0 0 0 |
| − | 13 51 1 0 0 0 0 | + | 13 51 1 0 0 0 0 |
| − | 51 52 1 0 0 0 0 | + | 51 52 1 0 0 0 0 |
| − | M STY 1 4 SUP | + | M STY 1 4 SUP |
| − | M SLB 1 4 4 | + | M SLB 1 4 4 |
| − | M SAL 4 2 47 48 | + | M SAL 4 2 47 48 |
| − | M SBL 4 1 51 | + | M SBL 4 1 51 |
| − | M SMT 4 CH2OH | + | M SMT 4 CH2OH |
| − | M SVB 4 51 3.9801 -1.2913 | + | M SVB 4 51 3.9801 -1.2913 |
| − | M STY 1 3 SUP | + | M STY 1 3 SUP |
| − | M SLB 1 3 3 | + | M SLB 1 3 3 |
| − | M SAL 3 2 51 52 | + | M SAL 3 2 51 52 |
| − | M SBL 3 1 55 | + | M SBL 3 1 55 |
| − | M SMT 3 OCH3 | + | M SMT 3 OCH3 |
| − | M SVB 3 55 2.7882 0.5751 | + | M SVB 3 55 2.7882 0.5751 |
| − | M STY 1 2 SUP | + | M STY 1 2 SUP |
| − | M SLB 1 2 2 | + | M SLB 1 2 2 |
| − | M SAL 2 2 49 50 | + | M SAL 2 2 49 50 |
| − | M SBL 2 1 53 | + | M SBL 2 1 53 |
| − | M SMT 2 OCH3 | + | M SMT 2 OCH3 |
| − | M SVB 2 53 2.295 2.3395 | + | M SVB 2 53 2.295 2.3395 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 3 43 44 46 | + | M SAL 1 3 43 44 46 |
| − | M SBL 1 1 47 | + | M SBL 1 1 47 |
| − | M SMT 1 COOH | + | M SMT 1 COOH |
| − | M SVB 1 47 -4.2101 -0.0854 | + | M SVB 1 47 -4.2101 -0.0854 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL7AAIGL0018 | + | ID FL7AAIGL0018 |
| − | KNApSAcK_ID C00006909 | + | KNApSAcK_ID C00006909 |
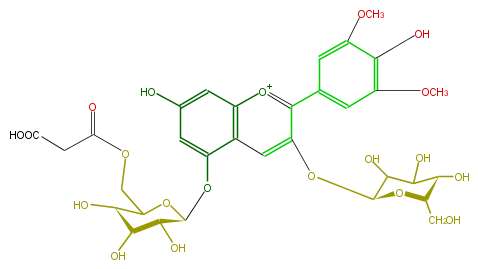
| − | NAME Malvidin 3-glucoside-5-(6''-malonylglucoside) | + | NAME Malvidin 3-glucoside-5-(6''-malonylglucoside) |
| − | CAS_RN 160206-02-0 | + | CAS_RN 160206-02-0 |
| − | FORMULA C32H37O20 | + | FORMULA C32H37O20 |
| − | EXACTMASS 741.187818624 | + | EXACTMASS 741.187818624 |
| − | AVERAGEMASS 741.62418 | + | AVERAGEMASS 741.62418 |
| − | SMILES O(c(c4c(c5)cc(OC)c(O)c(OC)5)cc(c([o+1]4)3)c(cc(O)c3)O[C@H](O2)[C@@H](O)[C@@H](O)[C@@H](O)C2COC(CC(O)=O)=O)[C@@H](C(O)1)O[C@@H]([C@@H](C1O)O)CO | + | SMILES O(c(c4c(c5)cc(OC)c(O)c(OC)5)cc(c([o+1]4)3)c(cc(O)c3)O[C@H](O2)[C@@H](O)[C@@H](O)[C@@H](O)C2COC(CC(O)=O)=O)[C@@H](C(O)1)O[C@@H]([C@@H](C1O)O)CO |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
52 56 0 0 0 0 0 0 0 0999 V2000
-1.3367 0.4603 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.3367 -0.1821 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.7804 -0.5032 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.2241 -0.1821 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.2241 0.4603 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.7804 0.7815 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.3322 -0.5032 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.8885 -0.1821 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.8885 0.4603 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.3322 0.7815 0.0000 O 0 3 0 0 0 0 0 0 0 0 0 0
1.4446 0.7814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0116 0.4540 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.5786 0.7814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.5786 1.4361 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0116 1.7634 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.4446 1.4361 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.8928 0.7814 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.4446 1.9361 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.7804 -1.1454 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.3046 -0.9028 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.8075 -0.9265 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
2.5393 -1.3911 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
3.0551 -1.2437 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.5740 -1.3911 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
3.8423 -0.9265 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
3.3264 -1.0739 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
2.4429 -0.5618 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.4681 -0.5451 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.2494 -0.9265 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.6504 -1.5433 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0
-2.2792 -2.0333 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
-1.7447 -1.8254 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
-1.2289 -1.8198 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
-1.6037 -1.4450 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.0713 -1.6918 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
-3.2413 -1.4916 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.6296 -2.4945 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.4384 -2.3395 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.5082 -1.1897 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.4206 -0.4639 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.0763 -0.0853 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.6433 -0.4126 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.2101 -0.0854 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.7770 -0.4126 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.0763 0.4818 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.2101 0.5691 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.7088 -1.7870 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.5655 -2.5994 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.2950 2.3395 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.7364 3.2368 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.5786 0.7814 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.5786 0.7814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 2 0 0 0 0
8 9 1 0 0 0 0
9 10 2 0 0 0 0
10 5 1 0 0 0 0
9 11 1 0 0 0 0
11 12 2 0 0 0 0
12 13 1 0 0 0 0
13 14 2 0 0 0 0
14 15 1 0 0 0 0
15 16 2 0 0 0 0
16 11 1 0 0 0 0
1 17 1 0 0 0 0
14 18 1 0 0 0 0
3 19 1 0 0 0 0
20 8 1 0 0 0 0
21 22 1 0 0 0 0
22 23 1 1 0 0 0
23 24 1 1 0 0 0
25 24 1 1 0 0 0
25 26 1 0 0 0 0
26 21 1 0 0 0 0
21 27 1 0 0 0 0
26 28 1 0 0 0 0
25 29 1 0 0 0 0
22 20 1 0 0 0 0
30 31 1 1 0 0 0
31 32 1 1 0 0 0
33 32 1 1 0 0 0
33 34 1 0 0 0 0
34 35 1 0 0 0 0
35 30 1 0 0 0 0
30 36 1 0 0 0 0
31 37 1 0 0 0 0
32 38 1 0 0 0 0
33 19 1 0 0 0 0
35 39 1 0 0 0 0
39 40 1 0 0 0 0
40 41 1 0 0 0 0
41 42 1 0 0 0 0
42 43 1 0 0 0 0
43 44 2 0 0 0 0
41 45 2 0 0 0 0
43 46 1 0 0 0 0
24 47 1 0 0 0 0
47 48 1 0 0 0 0
15 49 1 0 0 0 0
49 50 1 0 0 0 0
13 51 1 0 0 0 0
51 52 1 0 0 0 0
M STY 1 4 SUP
M SLB 1 4 4
M SAL 4 2 47 48
M SBL 4 1 51
M SMT 4 CH2OH
M SVB 4 51 3.9801 -1.2913
M STY 1 3 SUP
M SLB 1 3 3
M SAL 3 2 51 52
M SBL 3 1 55
M SMT 3 OCH3
M SVB 3 55 2.7882 0.5751
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 49 50
M SBL 2 1 53
M SMT 2 OCH3
M SVB 2 53 2.295 2.3395
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 3 43 44 46
M SBL 1 1 47
M SMT 1 COOH
M SVB 1 47 -4.2101 -0.0854
S SKP 8
ID FL7AAIGL0018
KNApSAcK_ID C00006909
NAME Malvidin 3-glucoside-5-(6''-malonylglucoside)
CAS_RN 160206-02-0
FORMULA C32H37O20
EXACTMASS 741.187818624
AVERAGEMASS 741.62418
SMILES O(c(c4c(c5)cc(OC)c(O)c(OC)5)cc(c([o+1]4)3)c(cc(O)c3)O[C@H](O2)[C@@H](O)[C@@H](O)[C@@H](O)C2COC(CC(O)=O)=O)[C@@H](C(O)1)O[C@@H]([C@@H](C1O)O)CO
M END