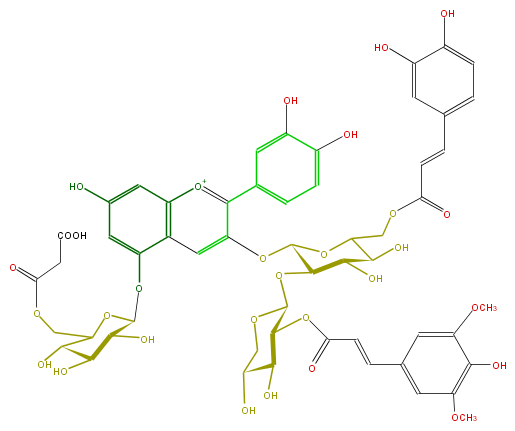
Mol:FL7AACGL0063
From Metabolomics.JP
Copyright: ARM project http://www.metabolome.jp/
85 92 0 0 0 0 0 0 0 0999 V2000
-3.6921 0.5448 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.6921 -0.2734 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.9834 -0.6826 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.2747 -0.2734 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.2747 0.5448 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.9834 0.9540 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.5660 -0.6826 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.8573 -0.2734 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.8573 0.5448 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.5660 0.9540 0.0000 O 0 3 0 0 0 0 0 0 0 0 0 0
-0.1489 0.9539 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5733 0.5369 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2956 0.9539 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2956 1.7879 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5733 2.2049 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.1489 1.7879 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.4005 0.9539 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.0177 2.2048 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.9834 -1.5006 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.0091 -0.7782 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.6811 -0.4631 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1928 -1.1386 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.9297 -0.8520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.6408 -0.8444 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.1241 -0.3275 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.4793 -0.6678 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.3951 -1.1774 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.6220 -1.2517 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.2402 -0.4983 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.5733 3.0387 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.8470 -2.9162 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.1254 -3.2204 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.6681 -2.6506 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.1028 -2.3159 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.7549 -2.1411 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.1137 -2.7113 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.3421 -3.3228 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.7844 -3.4989 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.8571 -2.7147 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-5.0271 -2.5824 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.4462 -2.0904 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.8759 -0.2420 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.1231 0.3047 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.8056 0.6573 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.4326 0.2953 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.8056 1.4723 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3558 1.7900 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3558 2.6559 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.0705 3.0685 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.0705 3.8939 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3558 4.3065 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.6411 3.8939 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.6411 3.0685 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3558 5.1315 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.4778 -3.5491 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.2638 -3.3577 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.2283 -2.6001 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5987 -1.9463 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.2147 -2.2333 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.1430 -3.0228 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.4338 -4.4222 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.9265 4.3064 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.0296 -2.1930 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.5664 -2.7503 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1675 -3.4134 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.2385 -2.7517 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.5547 -3.3418 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.3382 -3.3418 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.7508 -4.0565 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.5761 -4.0565 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.9887 -3.3418 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.5761 -2.6270 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.7508 -2.6270 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.6090 -3.3432 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-5.4462 -1.3104 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-6.0152 -0.9819 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.8818 -0.8883 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0972 -4.0728 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.1051 -1.9506 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
6.0152 -2.4761 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.6281 -4.6060 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.5383 -5.1315 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.8804 -0.2197 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.2985 -0.0638 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-5.4565 0.1129 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 2 0 0 0 0
8 9 1 0 0 0 0
9 10 2 0 0 0 0
10 5 1 0 0 0 0
9 11 1 0 0 0 0
11 12 2 0 0 0 0
12 13 1 0 0 0 0
13 14 2 0 0 0 0
14 15 1 0 0 0 0
15 16 2 0 0 0 0
16 11 1 0 0 0 0
1 17 1 0 0 0 0
14 18 1 0 0 0 0
3 19 1 0 0 0 0
20 8 1 0 0 0 0
21 22 1 1 0 0 0
22 23 1 1 0 0 0
24 23 1 1 0 0 0
24 25 1 0 0 0 0
25 26 1 0 0 0 0
26 21 1 0 0 0 0
22 27 1 0 0 0 0
23 28 1 0 0 0 0
24 29 1 0 0 0 0
15 30 1 0 0 0 0
31 32 1 1 0 0 0
32 33 1 1 0 0 0
34 33 1 1 0 0 0
34 35 1 0 0 0 0
35 36 1 0 0 0 0
36 31 1 0 0 0 0
31 37 1 0 0 0 0
32 38 1 0 0 0 0
33 39 1 0 0 0 0
36 40 1 0 0 0 0
40 41 1 0 0 0 0
34 19 1 0 0 0 0
25 42 1 0 0 0 0
42 43 1 0 0 0 0
44 45 2 0 0 0 0
44 46 1 0 0 0 0
46 47 2 0 0 0 0
47 48 1 0 0 0 0
44 43 1 0 0 0 0
48 49 2 0 0 0 0
49 50 1 0 0 0 0
50 51 2 0 0 0 0
51 52 1 0 0 0 0
52 53 2 0 0 0 0
53 48 1 0 0 0 0
51 54 1 0 0 0 0
55 56 1 1 0 0 0
56 57 1 1 0 0 0
58 57 1 1 0 0 0
58 59 1 0 0 0 0
59 60 1 0 0 0 0
60 55 1 0 0 0 0
55 61 1 0 0 0 0
20 21 1 0 0 0 0
27 58 1 0 0 0 0
52 62 1 0 0 0 0
64 65 2 0 0 0 0
64 66 1 0 0 0 0
66 67 2 0 0 0 0
67 68 1 0 0 0 0
64 63 1 0 0 0 0
68 69 2 0 0 0 0
69 70 1 0 0 0 0
70 71 2 0 0 0 0
71 72 1 0 0 0 0
72 73 2 0 0 0 0
73 68 1 0 0 0 0
71 74 1 0 0 0 0
63 57 1 0 0 0 0
41 75 1 0 0 0 0
75 76 2 0 0 0 0
75 77 1 0 0 0 0
56 78 1 0 0 0 0
79 80 1 0 0 0 0
72 79 1 0 0 0 0
81 82 1 0 0 0 0
70 81 1 0 0 0 0
83 85 1 0 0 0 0
83 84 2 0 0 0 0
77 83 1 0 0 0 0
M CHG 1 10 1
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 79 80
M SBL 1 1 87
M SMT 1 OCH3
M SBV 1 87 -0.5290 -0.6764
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 81 82
M SBL 2 1 89
M SMT 2 OCH3
M SBV 2 89 -0.0520 0.5496
M STY 1 3 SUP
M SLB 1 3 3
M SAL 3 3 83 84 85
M SBL 3 1 92
M SMT 3 COOH
M SBV 3 92 -0.0014 -0.6686
S SKP 5
ID FL7AACGL0063
FORMULA C55H57O30
EXACTMASS 1197.293465484
AVERAGEMASS 1198.02308
SMILES COc(c1O)cc(C=CC(OC(C(O)2)C(OC(C(O)3)C(Oc(c5)c(c(c8)cc(O)c(O)c8)[o+1]c(c7)c5c(cc(O)7)OC(O6)C(C(C(C(COC(CC(O)=O)=O)6)O)O)O)OC(COC(C=Cc(c4)cc(O)c(O)c4)=O)C3O)OCC2O)=O)cc1OC
M END