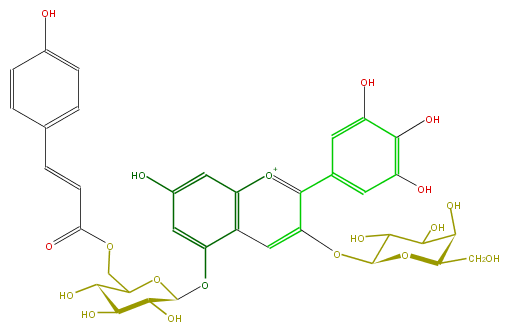
Mol:FL7AAGGA0007
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 55 60 0 0 0 0 0 0 0 0999 V2000 | + | 55 60 0 0 0 0 0 0 0 0999 V2000 |
| − | -1.9932 -0.6259 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.9932 -0.6259 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.9932 -1.4510 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.9932 -1.4510 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.2788 -1.8635 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.2788 -1.8635 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5643 -1.4510 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5643 -1.4510 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5643 -0.6259 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5643 -0.6259 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.2788 -0.2134 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.2788 -0.2134 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1502 -1.8635 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1502 -1.8635 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8646 -1.4510 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8646 -1.4510 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8646 -0.6259 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8646 -0.6259 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1502 -0.2134 0.0000 O 0 3 0 0 0 0 0 0 0 0 0 0 | + | 0.1502 -0.2134 0.0000 O 0 3 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5788 -0.2137 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5788 -0.2137 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.3070 -0.6340 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.3070 -0.6340 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0351 -0.2137 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0351 -0.2137 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0351 0.6272 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0351 0.6272 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.3070 1.0477 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.3070 1.0477 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5788 0.6272 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5788 0.6272 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7074 -0.2137 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7074 -0.2137 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.7632 1.0475 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.7632 1.0475 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.2788 -2.6881 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.2788 -2.6881 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.6878 -1.9847 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.6878 -1.9847 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.3070 1.8882 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.3070 1.8882 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.7213 -2.6897 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.7213 -2.6897 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.2446 -3.3188 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.2446 -3.3188 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.5581 -3.0520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.5581 -3.0520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.8958 -3.0448 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.8958 -3.0448 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.3770 -2.5634 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.3770 -2.5634 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.9776 -2.8804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.9776 -2.8804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.3179 -2.9071 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.3179 -2.9071 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.8213 -3.3358 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.8213 -3.3358 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.0602 -3.4455 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.0602 -3.4455 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.4509 -2.4418 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.4509 -2.4418 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.8768 -1.5613 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.8768 -1.5613 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.4905 -2.2304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.4905 -2.2304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.2335 -2.0181 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.2335 -2.0181 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.9808 -2.2304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.9808 -2.2304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.3673 -1.5613 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.3673 -1.5613 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.6243 -1.7735 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.6243 -1.7735 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2253 -1.6339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2253 -1.6339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.8697 -1.3895 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.8697 -1.3895 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.2270 -0.8793 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.2270 -0.8793 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.4303 -1.8108 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.4303 -1.8108 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.0985 -1.4251 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.0985 -1.4251 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.7898 -1.8243 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.7898 -1.8243 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.0985 -0.5139 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.0985 -0.5139 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.5898 -0.5338 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.5898 -0.5338 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.8848 -0.0599 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.8848 -0.0599 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.8848 0.8478 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.8848 0.8478 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.1347 1.2809 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.1347 1.2809 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.1347 2.1470 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.1347 2.1470 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.8848 2.5801 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.8848 2.5801 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.6349 2.1470 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.6349 2.1470 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.6349 1.2809 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.6349 1.2809 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.8848 3.4455 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.8848 3.4455 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.7174 -2.0836 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.7174 -2.0836 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.6349 -2.6134 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.6349 -2.6134 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 2 0 0 0 0 | + | 7 8 2 0 0 0 0 |
| − | 8 9 1 0 0 0 0 | + | 8 9 1 0 0 0 0 |
| − | 9 10 2 0 0 0 0 | + | 9 10 2 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 9 11 1 0 0 0 0 | + | 9 11 1 0 0 0 0 |
| − | 11 12 2 0 0 0 0 | + | 11 12 2 0 0 0 0 |
| − | 12 13 1 0 0 0 0 | + | 12 13 1 0 0 0 0 |
| − | 13 14 2 0 0 0 0 | + | 13 14 2 0 0 0 0 |
| − | 14 15 1 0 0 0 0 | + | 14 15 1 0 0 0 0 |
| − | 15 16 2 0 0 0 0 | + | 15 16 2 0 0 0 0 |
| − | 16 11 1 0 0 0 0 | + | 16 11 1 0 0 0 0 |
| − | 1 17 1 0 0 0 0 | + | 1 17 1 0 0 0 0 |
| − | 14 18 1 0 0 0 0 | + | 14 18 1 0 0 0 0 |
| − | 3 19 1 0 0 0 0 | + | 3 19 1 0 0 0 0 |
| − | 20 8 1 0 0 0 0 | + | 20 8 1 0 0 0 0 |
| − | 15 21 1 0 0 0 0 | + | 15 21 1 0 0 0 0 |
| − | 22 23 1 1 0 0 0 | + | 22 23 1 1 0 0 0 |
| − | 23 24 1 1 0 0 0 | + | 23 24 1 1 0 0 0 |
| − | 25 24 1 1 0 0 0 | + | 25 24 1 1 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 27 1 0 0 0 0 | + | 26 27 1 0 0 0 0 |
| − | 27 22 1 0 0 0 0 | + | 27 22 1 0 0 0 0 |
| − | 22 28 1 0 0 0 0 | + | 22 28 1 0 0 0 0 |
| − | 23 29 1 0 0 0 0 | + | 23 29 1 0 0 0 0 |
| − | 24 30 1 0 0 0 0 | + | 24 30 1 0 0 0 0 |
| − | 27 31 1 0 0 0 0 | + | 27 31 1 0 0 0 0 |
| − | 25 19 1 0 0 0 0 | + | 25 19 1 0 0 0 0 |
| − | 32 33 1 0 0 0 0 | + | 32 33 1 0 0 0 0 |
| − | 33 34 1 1 0 0 0 | + | 33 34 1 1 0 0 0 |
| − | 34 35 1 1 0 0 0 | + | 34 35 1 1 0 0 0 |
| − | 36 35 1 1 0 0 0 | + | 36 35 1 1 0 0 0 |
| − | 36 37 1 0 0 0 0 | + | 36 37 1 0 0 0 0 |
| − | 37 32 1 0 0 0 0 | + | 37 32 1 0 0 0 0 |
| − | 32 38 1 0 0 0 0 | + | 32 38 1 0 0 0 0 |
| − | 37 39 1 0 0 0 0 | + | 37 39 1 0 0 0 0 |
| − | 36 40 1 0 0 0 0 | + | 36 40 1 0 0 0 0 |
| − | 20 33 1 0 0 0 0 | + | 20 33 1 0 0 0 0 |
| − | 31 41 1 0 0 0 0 | + | 31 41 1 0 0 0 0 |
| − | 41 42 1 0 0 0 0 | + | 41 42 1 0 0 0 0 |
| − | 42 43 2 0 0 0 0 | + | 42 43 2 0 0 0 0 |
| − | 42 44 1 0 0 0 0 | + | 42 44 1 0 0 0 0 |
| − | 13 45 1 0 0 0 0 | + | 13 45 1 0 0 0 0 |
| − | 44 46 2 0 0 0 0 | + | 44 46 2 0 0 0 0 |
| − | 46 47 1 0 0 0 0 | + | 46 47 1 0 0 0 0 |
| − | 47 48 2 0 0 0 0 | + | 47 48 2 0 0 0 0 |
| − | 48 49 1 0 0 0 0 | + | 48 49 1 0 0 0 0 |
| − | 49 50 2 0 0 0 0 | + | 49 50 2 0 0 0 0 |
| − | 50 51 1 0 0 0 0 | + | 50 51 1 0 0 0 0 |
| − | 51 52 2 0 0 0 0 | + | 51 52 2 0 0 0 0 |
| − | 52 47 1 0 0 0 0 | + | 52 47 1 0 0 0 0 |
| − | 50 53 1 0 0 0 0 | + | 50 53 1 0 0 0 0 |
| − | 54 55 1 0 0 0 0 | + | 54 55 1 0 0 0 0 |
| − | 35 54 1 0 0 0 0 | + | 35 54 1 0 0 0 0 |
| − | M CHG 1 10 1 | + | M CHG 1 10 1 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 54 55 | + | M SAL 1 2 54 55 |
| − | M SBL 1 1 60 | + | M SBL 1 1 60 |
| − | M SMT 1 CH2OH | + | M SMT 1 CH2OH |
| − | M SBV 1 60 -0.7366 -0.1468 | + | M SBV 1 60 -0.7366 -0.1468 |
| − | S SKP 5 | + | S SKP 5 |
| − | ID FL7AAGGA0007 | + | ID FL7AAGGA0007 |
| − | FORMULA C36H37O19 | + | FORMULA C36H37O19 |
| − | EXACTMASS 773.192904002 | + | EXACTMASS 773.192904002 |
| − | AVERAGEMASS 773.66758 | + | AVERAGEMASS 773.66758 |
| − | SMILES OC(C6CO)C(C(O)C(O6)Oc(c5)c([o+1]c(c25)cc(cc2OC(C4O)OC(C(O)C4O)COC(C=Cc(c3)ccc(c3)O)=O)O)c(c1)cc(O)c(c1O)O)O | + | SMILES OC(C6CO)C(C(O)C(O6)Oc(c5)c([o+1]c(c25)cc(cc2OC(C4O)OC(C(O)C4O)COC(C=Cc(c3)ccc(c3)O)=O)O)c(c1)cc(O)c(c1O)O)O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
55 60 0 0 0 0 0 0 0 0999 V2000
-1.9932 -0.6259 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.9932 -1.4510 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.2788 -1.8635 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5643 -1.4510 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5643 -0.6259 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.2788 -0.2134 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.1502 -1.8635 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.8646 -1.4510 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.8646 -0.6259 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.1502 -0.2134 0.0000 O 0 3 0 0 0 0 0 0 0 0 0 0
1.5788 -0.2137 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.3070 -0.6340 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.0351 -0.2137 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.0351 0.6272 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.3070 1.0477 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.5788 0.6272 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7074 -0.2137 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.7632 1.0475 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.2788 -2.6881 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.6878 -1.9847 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.3070 1.8882 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.7213 -2.6897 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.2446 -3.3188 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.5581 -3.0520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.8958 -3.0448 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.3770 -2.5634 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.9776 -2.8804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.3179 -2.9071 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.8213 -3.3358 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.0602 -3.4455 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.4509 -2.4418 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.8768 -1.5613 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.4905 -2.2304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.2335 -2.0181 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.9808 -2.2304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3673 -1.5613 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.6243 -1.7735 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.2253 -1.6339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.8697 -1.3895 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.2270 -0.8793 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.4303 -1.8108 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.0985 -1.4251 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.7898 -1.8243 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.0985 -0.5139 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.5898 -0.5338 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.8848 -0.0599 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.8848 0.8478 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.1347 1.2809 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.1347 2.1470 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.8848 2.5801 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.6349 2.1470 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.6349 1.2809 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.8848 3.4455 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.7174 -2.0836 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.6349 -2.6134 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 2 0 0 0 0
8 9 1 0 0 0 0
9 10 2 0 0 0 0
10 5 1 0 0 0 0
9 11 1 0 0 0 0
11 12 2 0 0 0 0
12 13 1 0 0 0 0
13 14 2 0 0 0 0
14 15 1 0 0 0 0
15 16 2 0 0 0 0
16 11 1 0 0 0 0
1 17 1 0 0 0 0
14 18 1 0 0 0 0
3 19 1 0 0 0 0
20 8 1 0 0 0 0
15 21 1 0 0 0 0
22 23 1 1 0 0 0
23 24 1 1 0 0 0
25 24 1 1 0 0 0
25 26 1 0 0 0 0
26 27 1 0 0 0 0
27 22 1 0 0 0 0
22 28 1 0 0 0 0
23 29 1 0 0 0 0
24 30 1 0 0 0 0
27 31 1 0 0 0 0
25 19 1 0 0 0 0
32 33 1 0 0 0 0
33 34 1 1 0 0 0
34 35 1 1 0 0 0
36 35 1 1 0 0 0
36 37 1 0 0 0 0
37 32 1 0 0 0 0
32 38 1 0 0 0 0
37 39 1 0 0 0 0
36 40 1 0 0 0 0
20 33 1 0 0 0 0
31 41 1 0 0 0 0
41 42 1 0 0 0 0
42 43 2 0 0 0 0
42 44 1 0 0 0 0
13 45 1 0 0 0 0
44 46 2 0 0 0 0
46 47 1 0 0 0 0
47 48 2 0 0 0 0
48 49 1 0 0 0 0
49 50 2 0 0 0 0
50 51 1 0 0 0 0
51 52 2 0 0 0 0
52 47 1 0 0 0 0
50 53 1 0 0 0 0
54 55 1 0 0 0 0
35 54 1 0 0 0 0
M CHG 1 10 1
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 54 55
M SBL 1 1 60
M SMT 1 CH2OH
M SBV 1 60 -0.7366 -0.1468
S SKP 5
ID FL7AAGGA0007
FORMULA C36H37O19
EXACTMASS 773.192904002
AVERAGEMASS 773.66758
SMILES OC(C6CO)C(C(O)C(O6)Oc(c5)c([o+1]c(c25)cc(cc2OC(C4O)OC(C(O)C4O)COC(C=Cc(c3)ccc(c3)O)=O)O)c(c1)cc(O)c(c1O)O)O
M END