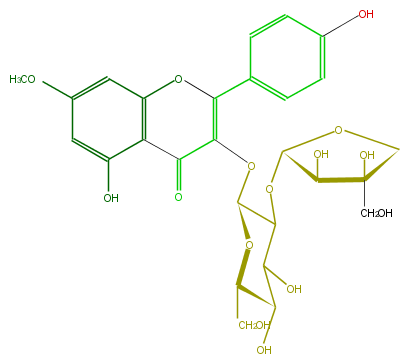
Mol:FL5FCAGL0002
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 42 46 0 0 0 0 0 0 0 0999 V2000 | + | 42 46 0 0 0 0 0 0 0 0999 V2000 |
| − | -2.7539 1.6446 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7539 1.6446 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7539 0.8255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7539 0.8255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.0444 0.4159 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.0444 0.4159 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.3350 0.8255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.3350 0.8255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.3350 1.6446 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.3350 1.6446 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.0444 2.0542 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.0444 2.0542 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6256 0.4159 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6256 0.4159 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0838 0.8255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0838 0.8255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0838 1.6446 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0838 1.6446 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6256 2.0542 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6256 2.0542 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6256 -0.3079 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6256 -0.3079 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7930 2.0541 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7930 2.0541 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5159 1.6366 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5159 1.6366 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2390 2.0541 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2390 2.0541 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2390 2.8890 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2390 2.8890 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5159 3.3064 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5159 3.3064 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7930 2.8890 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7930 2.8890 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8262 0.3114 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8262 0.3114 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.0444 -0.3289 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.0444 -0.3289 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0474 3.3557 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0474 3.3557 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2908 -0.8690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2908 -0.8690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5475 -0.4399 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5475 -0.4399 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7835 -1.2651 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7835 -1.2651 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5475 -2.0953 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5475 -2.0953 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2908 -2.5244 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2908 -2.5244 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.0550 -1.6992 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.0550 -1.6992 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1832 -0.1368 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1832 -0.1368 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.6036 -2.1435 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.6036 -2.1435 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.0191 -3.3557 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.0191 -3.3557 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.4466 0.5775 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.4466 0.5775 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.1453 0.0075 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.1453 0.0075 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0743 0.0227 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0743 0.0227 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.7755 0.6208 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.7755 0.6208 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.5793 1.0333 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5793 1.0333 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.1453 0.5604 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.1453 0.5604 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0743 0.5400 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0743 0.5400 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5342 -2.8703 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5342 -2.8703 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.7288 -3.0467 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7288 -3.0467 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.4529 2.0482 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.4529 2.0482 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.0905 3.1526 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.0905 3.1526 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0743 -0.6167 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0743 -0.6167 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.0905 -0.8890 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.0905 -0.8890 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 8 18 1 0 0 0 0 | + | 8 18 1 0 0 0 0 |
| − | 3 19 1 0 0 0 0 | + | 3 19 1 0 0 0 0 |
| − | 20 15 1 0 0 0 0 | + | 20 15 1 0 0 0 0 |
| − | 21 22 1 0 0 0 0 | + | 21 22 1 0 0 0 0 |
| − | 22 23 1 1 0 0 0 | + | 22 23 1 1 0 0 0 |
| − | 23 24 1 1 0 0 0 | + | 23 24 1 1 0 0 0 |
| − | 25 24 1 1 0 0 0 | + | 25 24 1 1 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 21 1 0 0 0 0 | + | 26 21 1 0 0 0 0 |
| − | 21 27 1 0 0 0 0 | + | 21 27 1 0 0 0 0 |
| − | 26 28 1 0 0 0 0 | + | 26 28 1 0 0 0 0 |
| − | 25 29 1 0 0 0 0 | + | 25 29 1 0 0 0 0 |
| − | 22 18 1 0 0 0 0 | + | 22 18 1 0 0 0 0 |
| − | 30 31 1 1 0 0 0 | + | 30 31 1 1 0 0 0 |
| − | 31 32 1 1 0 0 0 | + | 31 32 1 1 0 0 0 |
| − | 33 32 1 1 0 0 0 | + | 33 32 1 1 0 0 0 |
| − | 33 34 1 0 0 0 0 | + | 33 34 1 0 0 0 0 |
| − | 34 30 1 0 0 0 0 | + | 34 30 1 0 0 0 0 |
| − | 31 35 1 0 0 0 0 | + | 31 35 1 0 0 0 0 |
| − | 30 27 1 0 0 0 0 | + | 30 27 1 0 0 0 0 |
| − | 32 36 1 0 0 0 0 | + | 32 36 1 0 0 0 0 |
| − | 37 38 1 0 0 0 0 | + | 37 38 1 0 0 0 0 |
| − | 24 37 1 0 0 0 0 | + | 24 37 1 0 0 0 0 |
| − | 39 40 1 0 0 0 0 | + | 39 40 1 0 0 0 0 |
| − | 1 39 1 0 0 0 0 | + | 1 39 1 0 0 0 0 |
| − | 41 42 1 0 0 0 0 | + | 41 42 1 0 0 0 0 |
| − | 32 41 1 0 0 0 0 | + | 32 41 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 37 38 | + | M SAL 1 2 37 38 |
| − | M SBL 1 1 42 | + | M SBL 1 1 42 |
| − | M SMT 1 ^ CH2OH | + | M SMT 1 ^ CH2OH |
| − | M SBV 1 42 0.0133 0.7750 | + | M SBV 1 42 0.0133 0.7750 |
| − | M STY 1 2 SUP | + | M STY 1 2 SUP |
| − | M SLB 1 2 2 | + | M SLB 1 2 2 |
| − | M SAL 2 2 39 40 | + | M SAL 2 2 39 40 |
| − | M SBL 2 1 44 | + | M SBL 2 1 44 |
| − | M SMT 2 ^ OCH3 | + | M SMT 2 ^ OCH3 |
| − | M SBV 2 44 0.6991 -0.4036 | + | M SBV 2 44 0.6991 -0.4036 |
| − | M STY 1 3 SUP | + | M STY 1 3 SUP |
| − | M SLB 1 3 3 | + | M SLB 1 3 3 |
| − | M SAL 3 2 41 42 | + | M SAL 3 2 41 42 |
| − | M SBL 3 1 46 | + | M SBL 3 1 46 |
| − | M SMT 3 CH2OH | + | M SMT 3 CH2OH |
| − | M SBV 3 46 0.0000 0.6394 | + | M SBV 3 46 0.0000 0.6394 |
| − | S SKP 5 | + | S SKP 5 |
| − | ID FL5FCAGL0002 | + | ID FL5FCAGL0002 |
| − | FORMULA C27H30O15 | + | FORMULA C27H30O15 |
| − | EXACTMASS 594.15847029 | + | EXACTMASS 594.15847029 |
| − | AVERAGEMASS 594.5181 | + | AVERAGEMASS 594.5181 |
| − | SMILES C(OC(C3=O)=C(c(c5)ccc(c5)O)Oc(c4)c(c(O)cc4OC)3)(O2)C(C(O)C(C(CO)2)O)OC(C(O)1)OCC1(O)CO | + | SMILES C(OC(C3=O)=C(c(c5)ccc(c5)O)Oc(c4)c(c(O)cc4OC)3)(O2)C(C(O)C(C(CO)2)O)OC(C(O)1)OCC1(O)CO |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
42 46 0 0 0 0 0 0 0 0999 V2000
-2.7539 1.6446 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7539 0.8255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.0444 0.4159 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.3350 0.8255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.3350 1.6446 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.0444 2.0542 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.6256 0.4159 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0838 0.8255 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0838 1.6446 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.6256 2.0542 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.6256 -0.3079 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.7930 2.0541 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.5159 1.6366 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.2390 2.0541 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.2390 2.8890 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.5159 3.3064 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7930 2.8890 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.8262 0.3114 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.0444 -0.3289 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.0474 3.3557 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.2908 -0.8690 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5475 -0.4399 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7835 -1.2651 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.5475 -2.0953 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2908 -2.5244 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.0550 -1.6992 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1832 -0.1368 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.6036 -2.1435 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.0191 -3.3557 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.4466 0.5775 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.1453 0.0075 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.0743 0.0227 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.7755 0.6208 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.5793 1.0333 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.1453 0.5604 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.0743 0.5400 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.5342 -2.8703 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.7288 -3.0467 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.4529 2.0482 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.0905 3.1526 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.0743 -0.6167 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.0905 -0.8890 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
8 18 1 0 0 0 0
3 19 1 0 0 0 0
20 15 1 0 0 0 0
21 22 1 0 0 0 0
22 23 1 1 0 0 0
23 24 1 1 0 0 0
25 24 1 1 0 0 0
25 26 1 0 0 0 0
26 21 1 0 0 0 0
21 27 1 0 0 0 0
26 28 1 0 0 0 0
25 29 1 0 0 0 0
22 18 1 0 0 0 0
30 31 1 1 0 0 0
31 32 1 1 0 0 0
33 32 1 1 0 0 0
33 34 1 0 0 0 0
34 30 1 0 0 0 0
31 35 1 0 0 0 0
30 27 1 0 0 0 0
32 36 1 0 0 0 0
37 38 1 0 0 0 0
24 37 1 0 0 0 0
39 40 1 0 0 0 0
1 39 1 0 0 0 0
41 42 1 0 0 0 0
32 41 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 37 38
M SBL 1 1 42
M SMT 1 ^ CH2OH
M SBV 1 42 0.0133 0.7750
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 39 40
M SBL 2 1 44
M SMT 2 ^ OCH3
M SBV 2 44 0.6991 -0.4036
M STY 1 3 SUP
M SLB 1 3 3
M SAL 3 2 41 42
M SBL 3 1 46
M SMT 3 CH2OH
M SBV 3 46 0.0000 0.6394
S SKP 5
ID FL5FCAGL0002
FORMULA C27H30O15
EXACTMASS 594.15847029
AVERAGEMASS 594.5181
SMILES C(OC(C3=O)=C(c(c5)ccc(c5)O)Oc(c4)c(c(O)cc4OC)3)(O2)C(C(O)C(C(CO)2)O)OC(C(O)1)OCC1(O)CO
M END