Mol:FL5FAGGS0014
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 44 48 0 0 0 0 0 0 0 0999 V2000 | + | 44 48 0 0 0 0 0 0 0 0999 V2000 |
| − | -2.1977 1.7133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1977 1.7133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1977 1.0709 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1977 1.0709 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6414 0.7497 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6414 0.7497 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0851 1.0709 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0851 1.0709 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0851 1.7133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0851 1.7133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6414 2.0344 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6414 2.0344 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5288 0.7497 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5288 0.7497 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0275 1.0709 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0275 1.0709 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0275 1.7133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0275 1.7133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5288 2.0344 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5288 2.0344 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5288 0.2489 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5288 0.2489 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7280 2.1581 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7280 2.1581 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2950 1.8307 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2950 1.8307 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8619 2.1581 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8619 2.1581 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8619 2.8128 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8619 2.8128 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2950 3.1401 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2950 3.1401 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7280 2.8128 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7280 2.8128 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6414 0.1076 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6414 0.1076 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.4688 3.1631 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.4688 3.1631 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.6926 2.0344 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.6926 2.0344 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5908 0.6614 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5908 0.6614 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2950 3.7946 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2950 3.7946 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.6435 -0.3912 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.6435 -0.3912 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2718 -0.7539 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2718 -0.7539 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0724 -0.0564 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0724 -0.0564 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2718 0.6454 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2718 0.6454 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.6435 1.0081 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.6435 1.0081 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8428 0.3106 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8428 0.3106 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.6132 0.3106 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.6132 0.3106 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.6633 -1.3805 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.6633 -1.3805 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.1658 -1.1494 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.1658 -1.1494 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.6926 -0.4145 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.6926 -0.4145 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.4288 1.8308 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.4288 1.8308 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.6633 -2.0199 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.6633 -2.0199 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.1751 -2.3154 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.1751 -2.3154 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1759 -2.4470 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1759 -2.4470 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7091 -2.1774 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7091 -2.1774 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.2422 -2.4470 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.2422 -2.4470 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.2422 -2.9861 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.2422 -2.9861 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7091 -3.2556 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7091 -3.2556 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1759 -2.9861 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1759 -2.9861 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.2246 -2.1775 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.2246 -2.1775 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.2246 -3.2556 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.2246 -3.2556 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7091 -3.7946 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7091 -3.7946 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 3 18 1 0 0 0 0 | + | 3 18 1 0 0 0 0 |
| − | 19 15 1 0 0 0 0 | + | 19 15 1 0 0 0 0 |
| − | 20 1 1 0 0 0 0 | + | 20 1 1 0 0 0 0 |
| − | 21 8 1 0 0 0 0 | + | 21 8 1 0 0 0 0 |
| − | 4 3 1 0 0 0 0 | + | 4 3 1 0 0 0 0 |
| − | 16 22 1 0 0 0 0 | + | 16 22 1 0 0 0 0 |
| − | 24 23 1 1 0 0 0 | + | 24 23 1 1 0 0 0 |
| − | 24 25 1 0 0 0 0 | + | 24 25 1 0 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 27 1 0 0 0 0 | + | 26 27 1 0 0 0 0 |
| − | 27 28 1 1 0 0 0 | + | 27 28 1 1 0 0 0 |
| − | 28 23 1 1 0 0 0 | + | 28 23 1 1 0 0 0 |
| − | 28 29 1 0 0 0 0 | + | 28 29 1 0 0 0 0 |
| − | 23 30 1 0 0 0 0 | + | 23 30 1 0 0 0 0 |
| − | 24 31 1 0 0 0 0 | + | 24 31 1 0 0 0 0 |
| − | 25 32 1 0 0 0 0 | + | 25 32 1 0 0 0 0 |
| − | 27 21 1 0 0 0 0 | + | 27 21 1 0 0 0 0 |
| − | 14 33 1 0 0 0 0 | + | 14 33 1 0 0 0 0 |
| − | 30 34 1 0 0 0 0 | + | 30 34 1 0 0 0 0 |
| − | 34 35 2 0 0 0 0 | + | 34 35 2 0 0 0 0 |
| − | 34 36 1 0 0 0 0 | + | 34 36 1 0 0 0 0 |
| − | 36 37 2 0 0 0 0 | + | 36 37 2 0 0 0 0 |
| − | 37 38 1 0 0 0 0 | + | 37 38 1 0 0 0 0 |
| − | 38 39 2 0 0 0 0 | + | 38 39 2 0 0 0 0 |
| − | 39 40 1 0 0 0 0 | + | 39 40 1 0 0 0 0 |
| − | 40 41 2 0 0 0 0 | + | 40 41 2 0 0 0 0 |
| − | 41 36 1 0 0 0 0 | + | 41 36 1 0 0 0 0 |
| − | 38 42 1 0 0 0 0 | + | 38 42 1 0 0 0 0 |
| − | 39 43 1 0 0 0 0 | + | 39 43 1 0 0 0 0 |
| − | 40 44 1 0 0 0 0 | + | 40 44 1 0 0 0 0 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL5FAGGS0014 | + | ID FL5FAGGS0014 |
| − | KNApSAcK_ID C00006043 | + | KNApSAcK_ID C00006043 |
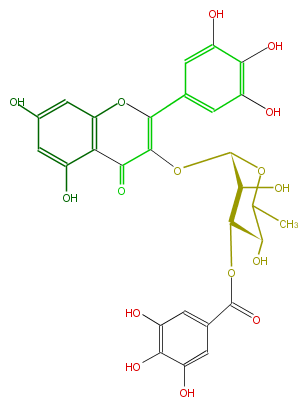
| − | NAME Myricetin 3-(3''-galloylrhamnoside) | + | NAME Myricetin 3-(3''-galloylrhamnoside) |
| − | CAS_RN 143202-36-2 | + | CAS_RN 143202-36-2 |
| − | FORMULA C28H24O16 | + | FORMULA C28H24O16 |
| − | EXACTMASS 616.1064347199999 | + | EXACTMASS 616.1064347199999 |
| − | AVERAGEMASS 616.48056 | + | AVERAGEMASS 616.48056 |
| − | SMILES c(O)(c5O)c(cc(c5)C(=C(OC(C(O)3)OC(C(C3OC(c(c4)cc(O)c(O)c(O)4)=O)O)C)2)Oc(c1C(=O)2)cc(cc(O)1)O)O | + | SMILES c(O)(c5O)c(cc(c5)C(=C(OC(C(O)3)OC(C(C3OC(c(c4)cc(O)c(O)c(O)4)=O)O)C)2)Oc(c1C(=O)2)cc(cc(O)1)O)O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
44 48 0 0 0 0 0 0 0 0999 V2000
-2.1977 1.7133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.1977 1.0709 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6414 0.7497 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.0851 1.0709 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.0851 1.7133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6414 2.0344 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5288 0.7497 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0275 1.0709 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0275 1.7133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5288 2.0344 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.5288 0.2489 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.7280 2.1581 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2950 1.8307 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8619 2.1581 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8619 2.8128 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2950 3.1401 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7280 2.8128 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6414 0.1076 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.4688 3.1631 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.6926 2.0344 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.5908 0.6614 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.2950 3.7946 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.6435 -0.3912 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.2718 -0.7539 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0724 -0.0564 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.2718 0.6454 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.6435 1.0081 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8428 0.3106 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.6132 0.3106 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.6633 -1.3805 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.1658 -1.1494 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.6926 -0.4145 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.4288 1.8308 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.6633 -2.0199 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.1751 -2.3154 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.1759 -2.4470 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7091 -2.1774 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.2422 -2.4470 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.2422 -2.9861 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7091 -3.2556 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1759 -2.9861 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.2246 -2.1775 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.2246 -3.2556 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.7091 -3.7946 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
3 18 1 0 0 0 0
19 15 1 0 0 0 0
20 1 1 0 0 0 0
21 8 1 0 0 0 0
4 3 1 0 0 0 0
16 22 1 0 0 0 0
24 23 1 1 0 0 0
24 25 1 0 0 0 0
25 26 1 0 0 0 0
26 27 1 0 0 0 0
27 28 1 1 0 0 0
28 23 1 1 0 0 0
28 29 1 0 0 0 0
23 30 1 0 0 0 0
24 31 1 0 0 0 0
25 32 1 0 0 0 0
27 21 1 0 0 0 0
14 33 1 0 0 0 0
30 34 1 0 0 0 0
34 35 2 0 0 0 0
34 36 1 0 0 0 0
36 37 2 0 0 0 0
37 38 1 0 0 0 0
38 39 2 0 0 0 0
39 40 1 0 0 0 0
40 41 2 0 0 0 0
41 36 1 0 0 0 0
38 42 1 0 0 0 0
39 43 1 0 0 0 0
40 44 1 0 0 0 0
S SKP 8
ID FL5FAGGS0014
KNApSAcK_ID C00006043
NAME Myricetin 3-(3''-galloylrhamnoside)
CAS_RN 143202-36-2
FORMULA C28H24O16
EXACTMASS 616.1064347199999
AVERAGEMASS 616.48056
SMILES c(O)(c5O)c(cc(c5)C(=C(OC(C(O)3)OC(C(C3OC(c(c4)cc(O)c(O)c(O)4)=O)O)C)2)Oc(c1C(=O)2)cc(cc(O)1)O)O
M END