Mol:FL5FADGL0026
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 56 61 0 0 0 0 0 0 0 0999 V2000 | + | 56 61 0 0 0 0 0 0 0 0999 V2000 |
| − | 1.4254 -4.5718 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.4254 -4.5718 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0579 -4.4602 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0579 -4.4602 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2776 -3.8566 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2776 -3.8566 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8647 -3.3645 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8647 -3.3645 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2322 -3.4761 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2322 -3.4761 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.0124 -4.0797 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.0124 -4.0797 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0844 -2.7609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0844 -2.7609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.6715 -2.2688 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.6715 -2.2688 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.0390 -2.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.0390 -2.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8192 -2.9840 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8192 -2.9840 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.5777 -2.6739 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5777 -2.6739 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6262 -1.8885 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6262 -1.8885 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8501 -1.2732 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8501 -1.2732 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.4293 -0.7718 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.4293 -0.7718 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.2155 -0.8855 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.2155 -0.8855 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.4394 -1.5006 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.4394 -1.5006 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0186 -2.0022 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0186 -2.0022 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2057 -5.1752 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2057 -5.1752 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.9099 -3.7450 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.9099 -3.7450 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.8583 -0.1195 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.8583 -0.1195 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.6762 -0.2803 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 2.6762 -0.2803 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 2.3630 -0.8228 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.3630 -0.8228 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 2.9653 -0.6507 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.9653 -0.6507 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.5713 -0.8228 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 3.5713 -0.8228 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 3.8846 -0.2803 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 3.8846 -0.2803 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 3.2822 -0.4524 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 3.2822 -0.4524 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 2.1343 -0.2803 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.1343 -0.2803 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.3193 -0.0288 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.3193 -0.0288 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.5626 -0.4620 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.5626 -0.4620 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.0918 -0.5801 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.0918 -0.5801 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 6.4641 0.0707 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 | + | 6.4641 0.0707 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 |
| − | 6.0879 0.6694 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 6.0879 0.6694 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 5.4546 0.4848 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 5.4546 0.4848 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 4.8634 0.5302 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 4.8634 0.5302 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 5.2549 0.0630 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.2549 0.0630 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.9002 0.2329 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 5.9002 0.2329 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 7.1267 -0.0461 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 7.1267 -0.0461 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 6.5356 1.1629 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 6.5356 1.1629 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.1555 1.1046 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.1555 1.1046 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.4075 -0.1207 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.4075 -0.1207 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.1503 -1.5507 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.1503 -1.5507 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0459 -7.4265 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 | + | 2.0459 -7.4265 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 |
| − | 2.5267 -6.9081 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.5267 -6.9081 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 2.1845 -6.3442 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.1845 -6.3442 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 2.0752 -5.7611 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.0752 -5.7611 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 1.7255 -6.2605 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.7255 -6.2605 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0566 -6.8398 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 2.0566 -6.8398 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 2.0940 -7.9755 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0940 -7.9755 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.1193 -7.2129 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.1193 -7.2129 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.7057 -5.8946 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.7057 -5.8946 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.8757 -0.5765 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.8757 -0.5765 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 6.2708 -1.4952 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 6.2708 -1.4952 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0559 -1.3216 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0559 -1.3216 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.0163 -1.0427 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.0163 -1.0427 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3827 -7.1975 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3827 -7.1975 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3470 -8.1968 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3470 -8.1968 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 1 18 1 0 0 0 0 | + | 1 18 1 0 0 0 0 |
| − | 3 19 1 0 0 0 0 | + | 3 19 1 0 0 0 0 |
| − | 20 15 1 0 0 0 0 | + | 20 15 1 0 0 0 0 |
| − | 21 22 1 0 0 0 0 | + | 21 22 1 0 0 0 0 |
| − | 22 23 1 1 0 0 0 | + | 22 23 1 1 0 0 0 |
| − | 23 24 1 1 0 0 0 | + | 23 24 1 1 0 0 0 |
| − | 25 24 1 1 0 0 0 | + | 25 24 1 1 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 21 1 0 0 0 0 | + | 26 21 1 0 0 0 0 |
| − | 21 27 1 0 0 0 0 | + | 21 27 1 0 0 0 0 |
| − | 26 28 1 0 0 0 0 | + | 26 28 1 0 0 0 0 |
| − | 25 29 1 0 0 0 0 | + | 25 29 1 0 0 0 0 |
| − | 24 30 1 0 0 0 0 | + | 24 30 1 0 0 0 0 |
| − | 31 32 1 1 0 0 0 | + | 31 32 1 1 0 0 0 |
| − | 32 33 1 1 0 0 0 | + | 32 33 1 1 0 0 0 |
| − | 34 33 1 1 0 0 0 | + | 34 33 1 1 0 0 0 |
| − | 34 35 1 0 0 0 0 | + | 34 35 1 0 0 0 0 |
| − | 35 36 1 0 0 0 0 | + | 35 36 1 0 0 0 0 |
| − | 36 31 1 0 0 0 0 | + | 36 31 1 0 0 0 0 |
| − | 31 37 1 0 0 0 0 | + | 31 37 1 0 0 0 0 |
| − | 32 38 1 0 0 0 0 | + | 32 38 1 0 0 0 0 |
| − | 33 39 1 0 0 0 0 | + | 33 39 1 0 0 0 0 |
| − | 34 40 1 0 0 0 0 | + | 34 40 1 0 0 0 0 |
| − | 40 30 1 0 0 0 0 | + | 40 30 1 0 0 0 0 |
| − | 22 41 1 0 0 0 0 | + | 22 41 1 0 0 0 0 |
| − | 41 8 1 0 0 0 0 | + | 41 8 1 0 0 0 0 |
| − | 42 43 1 1 0 0 0 | + | 42 43 1 1 0 0 0 |
| − | 43 44 1 1 0 0 0 | + | 43 44 1 1 0 0 0 |
| − | 45 44 1 1 0 0 0 | + | 45 44 1 1 0 0 0 |
| − | 45 46 1 0 0 0 0 | + | 45 46 1 0 0 0 0 |
| − | 46 47 1 0 0 0 0 | + | 46 47 1 0 0 0 0 |
| − | 47 42 1 0 0 0 0 | + | 47 42 1 0 0 0 0 |
| − | 42 48 1 0 0 0 0 | + | 42 48 1 0 0 0 0 |
| − | 43 49 1 0 0 0 0 | + | 43 49 1 0 0 0 0 |
| − | 44 50 1 0 0 0 0 | + | 44 50 1 0 0 0 0 |
| − | 45 18 1 0 0 0 0 | + | 45 18 1 0 0 0 0 |
| − | 36 51 1 0 0 0 0 | + | 36 51 1 0 0 0 0 |
| − | 51 52 1 0 0 0 0 | + | 51 52 1 0 0 0 0 |
| − | 16 53 1 0 0 0 0 | + | 16 53 1 0 0 0 0 |
| − | 53 54 1 0 0 0 0 | + | 53 54 1 0 0 0 0 |
| − | 47 55 1 0 0 0 0 | + | 47 55 1 0 0 0 0 |
| − | 55 56 1 0 0 0 0 | + | 55 56 1 0 0 0 0 |
| − | M STY 1 3 SUP | + | M STY 1 3 SUP |
| − | M SLB 1 3 3 | + | M SLB 1 3 3 |
| − | M SAL 3 2 55 56 | + | M SAL 3 2 55 56 |
| − | M SBL 3 1 60 | + | M SBL 3 1 60 |
| − | M SMT 3 CH2OH | + | M SMT 3 CH2OH |
| − | M SVB 3 60 -3.7446 1.2496 | + | M SVB 3 60 -3.7446 1.2496 |
| − | M STY 1 2 SUP | + | M STY 1 2 SUP |
| − | M SLB 1 2 2 | + | M SLB 1 2 2 |
| − | M SAL 2 2 51 52 | + | M SAL 2 2 51 52 |
| − | M SBL 2 1 56 | + | M SBL 2 1 56 |
| − | M SMT 2 CH2OH | + | M SMT 2 CH2OH |
| − | M SVB 2 56 2.6986 -1.0393 | + | M SVB 2 56 2.6986 -1.0393 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 53 54 | + | M SAL 1 2 53 54 |
| − | M SBL 1 1 58 | + | M SBL 1 1 58 |
| − | M SMT 1 OCH3 | + | M SMT 1 OCH3 |
| − | M SVB 1 58 2.4655 2.6308 | + | M SVB 1 58 2.4655 2.6308 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL5FADGL0026 | + | ID FL5FADGL0026 |
| − | KNApSAcK_ID C00005581 | + | KNApSAcK_ID C00005581 |
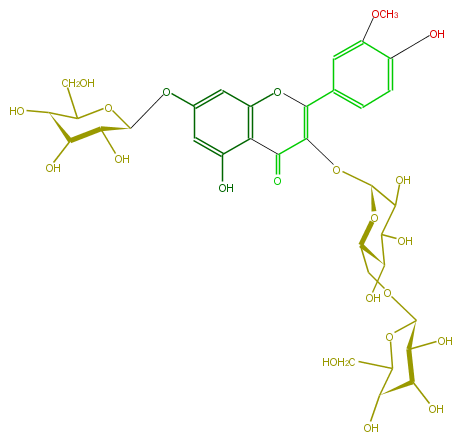
| − | NAME Isorhamnetin 3-gentiobioside-7-glucoside | + | NAME Isorhamnetin 3-gentiobioside-7-glucoside |
| − | CAS_RN - | + | CAS_RN - |
| − | FORMULA C34H42O22 | + | FORMULA C34H42O22 |
| − | EXACTMASS 802.216773028 | + | EXACTMASS 802.216773028 |
| − | AVERAGEMASS 802.68408 | + | AVERAGEMASS 802.68408 |
| − | SMILES O[C@H]([C@@H]1OC[C@@H](O2)[C@@H](C(C(O)[C@H](OC(=C(c(c6)cc(c(c6)O)OC)3)C(c(c4O)c(cc(O[C@@H]([C@H]5O)OC([C@@H]([C@H](O)5)O)CO)c4)O3)=O)2)O)O)[C@@H](O)[C@@H](O)C(O1)CO | + | SMILES O[C@H]([C@@H]1OC[C@@H](O2)[C@@H](C(C(O)[C@H](OC(=C(c(c6)cc(c(c6)O)OC)3)C(c(c4O)c(cc(O[C@@H]([C@H]5O)OC([C@@H]([C@H](O)5)O)CO)c4)O3)=O)2)O)O)[C@@H](O)[C@@H](O)C(O1)CO |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
56 61 0 0 0 0 0 0 0 0999 V2000
1.4254 -4.5718 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0579 -4.4602 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.2776 -3.8566 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8647 -3.3645 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2322 -3.4761 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.0124 -4.0797 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0844 -2.7609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.6715 -2.2688 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.0390 -2.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.8192 -2.9840 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.5777 -2.6739 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.6262 -1.8885 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.8501 -1.2732 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.4293 -0.7718 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.2155 -0.8855 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.4394 -1.5006 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.0186 -2.0022 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2057 -5.1752 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.9099 -3.7450 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.8583 -0.1195 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.6762 -0.2803 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
2.3630 -0.8228 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
2.9653 -0.6507 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.5713 -0.8228 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
3.8846 -0.2803 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
3.2822 -0.4524 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
2.1343 -0.2803 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.3193 -0.0288 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.5626 -0.4620 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.0918 -0.5801 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
6.4641 0.0707 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0
6.0879 0.6694 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
5.4546 0.4848 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
4.8634 0.5302 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
5.2549 0.0630 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.9002 0.2329 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
7.1267 -0.0461 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
6.5356 1.1629 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.1555 1.1046 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.4075 -0.1207 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.1503 -1.5507 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.0459 -7.4265 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0
2.5267 -6.9081 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
2.1845 -6.3442 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
2.0752 -5.7611 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
1.7255 -6.2605 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.0566 -6.8398 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
2.0940 -7.9755 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.1193 -7.2129 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.7057 -5.8946 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.8757 -0.5765 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
6.2708 -1.4952 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.0559 -1.3216 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.0163 -1.0427 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.3827 -7.1975 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.3470 -8.1968 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
1 18 1 0 0 0 0
3 19 1 0 0 0 0
20 15 1 0 0 0 0
21 22 1 0 0 0 0
22 23 1 1 0 0 0
23 24 1 1 0 0 0
25 24 1 1 0 0 0
25 26 1 0 0 0 0
26 21 1 0 0 0 0
21 27 1 0 0 0 0
26 28 1 0 0 0 0
25 29 1 0 0 0 0
24 30 1 0 0 0 0
31 32 1 1 0 0 0
32 33 1 1 0 0 0
34 33 1 1 0 0 0
34 35 1 0 0 0 0
35 36 1 0 0 0 0
36 31 1 0 0 0 0
31 37 1 0 0 0 0
32 38 1 0 0 0 0
33 39 1 0 0 0 0
34 40 1 0 0 0 0
40 30 1 0 0 0 0
22 41 1 0 0 0 0
41 8 1 0 0 0 0
42 43 1 1 0 0 0
43 44 1 1 0 0 0
45 44 1 1 0 0 0
45 46 1 0 0 0 0
46 47 1 0 0 0 0
47 42 1 0 0 0 0
42 48 1 0 0 0 0
43 49 1 0 0 0 0
44 50 1 0 0 0 0
45 18 1 0 0 0 0
36 51 1 0 0 0 0
51 52 1 0 0 0 0
16 53 1 0 0 0 0
53 54 1 0 0 0 0
47 55 1 0 0 0 0
55 56 1 0 0 0 0
M STY 1 3 SUP
M SLB 1 3 3
M SAL 3 2 55 56
M SBL 3 1 60
M SMT 3 CH2OH
M SVB 3 60 -3.7446 1.2496
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 51 52
M SBL 2 1 56
M SMT 2 CH2OH
M SVB 2 56 2.6986 -1.0393
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 53 54
M SBL 1 1 58
M SMT 1 OCH3
M SVB 1 58 2.4655 2.6308
S SKP 8
ID FL5FADGL0026
KNApSAcK_ID C00005581
NAME Isorhamnetin 3-gentiobioside-7-glucoside
CAS_RN -
FORMULA C34H42O22
EXACTMASS 802.216773028
AVERAGEMASS 802.68408
SMILES O[C@H]([C@@H]1OC[C@@H](O2)[C@@H](C(C(O)[C@H](OC(=C(c(c6)cc(c(c6)O)OC)3)C(c(c4O)c(cc(O[C@@H]([C@H]5O)OC([C@@H]([C@H](O)5)O)CO)c4)O3)=O)2)O)O)[C@@H](O)[C@@H](O)C(O1)CO
M END