Mol:FL3FADGS0009
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 43 47 0 0 0 0 0 0 0 0999 V2000 | + | 43 47 0 0 0 0 0 0 0 0999 V2000 |
| − | 0.6173 -0.9757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6173 -0.9757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6173 -1.7579 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6173 -1.7579 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2946 -2.1489 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2946 -2.1489 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.9719 -1.7579 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.9719 -1.7579 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.9719 -0.9757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.9719 -0.9757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2946 -0.5847 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2946 -0.5847 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.6493 -2.1489 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.6493 -2.1489 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.3265 -1.7579 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.3265 -1.7579 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.3265 -0.9757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.3265 -0.9757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.6493 -0.5847 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.6493 -0.5847 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.9071 -2.7013 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.9071 -2.7013 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.0036 -0.5849 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.0036 -0.5849 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.6938 -0.9834 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.6938 -0.9834 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.3843 -0.5849 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.3843 -0.5849 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.3843 0.2122 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.3843 0.2122 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.6938 0.6107 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.6938 0.6107 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.0036 0.2122 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.0036 0.2122 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2946 -2.9294 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2946 -2.9294 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 6.0291 0.6126 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 6.0291 0.6126 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0587 -0.5856 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0587 -0.5856 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.4741 -0.3385 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.4741 -0.3385 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7773 -1.2583 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7773 -1.2583 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.7739 -0.8682 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.7739 -0.8682 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.8056 -0.8578 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.8056 -0.8578 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.5093 -0.1539 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.5093 -0.1539 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.3871 -0.6173 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.3871 -0.6173 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.8925 -0.0651 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.8925 -0.0651 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.2311 -0.7318 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.2311 -0.7318 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.4909 -1.2714 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.4909 -1.2714 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.8676 -1.4635 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.8676 -1.4635 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.3545 0.3351 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.3545 0.3351 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.5054 -0.1551 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.5054 -0.1551 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.7749 0.7876 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.7749 0.7876 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.5054 1.7359 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.5054 1.7359 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.3545 2.2261 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.3545 2.2261 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.0851 1.2834 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.0851 1.2834 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.6074 -0.0280 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.6074 -0.0280 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.0761 2.9294 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.0761 2.9294 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.5105 2.4528 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.5105 2.4528 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.6825 1.7961 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.6825 1.7961 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -6.0291 0.2859 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -6.0291 0.2859 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.7104 1.3455 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.7104 1.3455 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.2938 2.7290 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.2938 2.7290 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 3 18 1 0 0 0 0 | + | 3 18 1 0 0 0 0 |
| − | 19 15 1 0 0 0 0 | + | 19 15 1 0 0 0 0 |
| − | 1 20 1 0 0 0 0 | + | 1 20 1 0 0 0 0 |
| − | 21 22 1 1 0 0 0 | + | 21 22 1 1 0 0 0 |
| − | 22 23 1 1 0 0 0 | + | 22 23 1 1 0 0 0 |
| − | 24 23 1 1 0 0 0 | + | 24 23 1 1 0 0 0 |
| − | 24 25 1 0 0 0 0 | + | 24 25 1 0 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 21 1 0 0 0 0 | + | 26 21 1 0 0 0 0 |
| − | 26 27 1 0 0 0 0 | + | 26 27 1 0 0 0 0 |
| − | 21 28 1 0 0 0 0 | + | 21 28 1 0 0 0 0 |
| − | 22 29 1 0 0 0 0 | + | 22 29 1 0 0 0 0 |
| − | 23 30 1 0 0 0 0 | + | 23 30 1 0 0 0 0 |
| − | 32 31 1 1 0 0 0 | + | 32 31 1 1 0 0 0 |
| − | 32 33 1 0 0 0 0 | + | 32 33 1 0 0 0 0 |
| − | 33 34 1 0 0 0 0 | + | 33 34 1 0 0 0 0 |
| − | 34 35 1 0 0 0 0 | + | 34 35 1 0 0 0 0 |
| − | 35 36 1 1 0 0 0 | + | 35 36 1 1 0 0 0 |
| − | 36 31 1 1 0 0 0 | + | 36 31 1 1 0 0 0 |
| − | 32 37 1 0 0 0 0 | + | 32 37 1 0 0 0 0 |
| − | 37 27 1 0 0 0 0 | + | 37 27 1 0 0 0 0 |
| − | 35 38 1 0 0 0 0 | + | 35 38 1 0 0 0 0 |
| − | 34 39 1 0 0 0 0 | + | 34 39 1 0 0 0 0 |
| − | 36 40 1 0 0 0 0 | + | 36 40 1 0 0 0 0 |
| − | 31 41 1 0 0 0 0 | + | 31 41 1 0 0 0 0 |
| − | 24 20 1 0 0 0 0 | + | 24 20 1 0 0 0 0 |
| − | 42 43 1 0 0 0 0 | + | 42 43 1 0 0 0 0 |
| − | 16 42 1 0 0 0 0 | + | 16 42 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 42 43 | + | M SAL 1 2 42 43 |
| − | M SBL 1 1 47 | + | M SBL 1 1 47 |
| − | M SMT 1 OCH3 | + | M SMT 1 OCH3 |
| − | M SBV 1 47 -0.0166 -0.7348 | + | M SBV 1 47 -0.0166 -0.7348 |
| − | S SKP 5 | + | S SKP 5 |
| − | ID FL3FADGS0009 | + | ID FL3FADGS0009 |
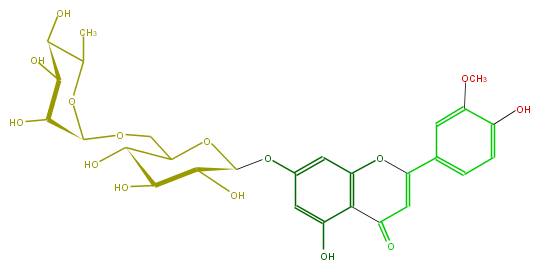
| − | FORMULA C28H32O15 | + | FORMULA C28H32O15 |
| − | EXACTMASS 608.174120354 | + | EXACTMASS 608.174120354 |
| − | AVERAGEMASS 608.54468 | + | AVERAGEMASS 608.54468 |
| − | SMILES OC(C(O)4)C(COC(O5)C(C(C(O)C(C)5)O)O)OC(C(O)4)Oc(c3)cc(c(c23)C(=O)C=C(O2)c(c1)ccc(O)c(OC)1)O | + | SMILES OC(C(O)4)C(COC(O5)C(C(C(O)C(C)5)O)O)OC(C(O)4)Oc(c3)cc(c(c23)C(=O)C=C(O2)c(c1)ccc(O)c(OC)1)O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
43 47 0 0 0 0 0 0 0 0999 V2000
0.6173 -0.9757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.6173 -1.7579 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2946 -2.1489 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.9719 -1.7579 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.9719 -0.9757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2946 -0.5847 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.6493 -2.1489 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.3265 -1.7579 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.3265 -0.9757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.6493 -0.5847 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.9071 -2.7013 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.0036 -0.5849 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.6938 -0.9834 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.3843 -0.5849 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.3843 0.2122 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.6938 0.6107 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.0036 0.2122 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2946 -2.9294 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
6.0291 0.6126 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.0587 -0.5856 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.4741 -0.3385 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7773 -1.2583 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.7739 -0.8682 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.8056 -0.8578 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.5093 -0.1539 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.3871 -0.6173 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.8925 -0.0651 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.2311 -0.7318 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.4909 -1.2714 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.8676 -1.4635 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-5.3545 0.3351 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.5054 -0.1551 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.7749 0.7876 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.5054 1.7359 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.3545 2.2261 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.0851 1.2834 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.6074 -0.0280 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-5.0761 2.9294 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.5105 2.4528 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.6825 1.7961 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-6.0291 0.2859 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.7104 1.3455 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.2938 2.7290 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
3 18 1 0 0 0 0
19 15 1 0 0 0 0
1 20 1 0 0 0 0
21 22 1 1 0 0 0
22 23 1 1 0 0 0
24 23 1 1 0 0 0
24 25 1 0 0 0 0
25 26 1 0 0 0 0
26 21 1 0 0 0 0
26 27 1 0 0 0 0
21 28 1 0 0 0 0
22 29 1 0 0 0 0
23 30 1 0 0 0 0
32 31 1 1 0 0 0
32 33 1 0 0 0 0
33 34 1 0 0 0 0
34 35 1 0 0 0 0
35 36 1 1 0 0 0
36 31 1 1 0 0 0
32 37 1 0 0 0 0
37 27 1 0 0 0 0
35 38 1 0 0 0 0
34 39 1 0 0 0 0
36 40 1 0 0 0 0
31 41 1 0 0 0 0
24 20 1 0 0 0 0
42 43 1 0 0 0 0
16 42 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 42 43
M SBL 1 1 47
M SMT 1 OCH3
M SBV 1 47 -0.0166 -0.7348
S SKP 5
ID FL3FADGS0009
FORMULA C28H32O15
EXACTMASS 608.174120354
AVERAGEMASS 608.54468
SMILES OC(C(O)4)C(COC(O5)C(C(C(O)C(C)5)O)O)OC(C(O)4)Oc(c3)cc(c(c23)C(=O)C=C(O2)c(c1)ccc(O)c(OC)1)O
M END