Mol:FL3FACGS0056
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 42 46 0 0 0 0 0 0 0 0999 V2000 | + | 42 46 0 0 0 0 0 0 0 0999 V2000 |
| − | -0.5298 -0.9377 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5298 -0.9377 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5298 -1.7473 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5298 -1.7473 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1714 -2.1520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1714 -2.1520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8723 -1.7473 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8723 -1.7473 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8723 -0.9377 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8723 -0.9377 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1714 -0.5330 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1714 -0.5330 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5734 -2.1520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5734 -2.1520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2745 -1.7473 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2745 -1.7473 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2745 -0.9377 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2745 -0.9377 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5734 -0.5330 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5734 -0.5330 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5734 -2.9803 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5734 -2.9803 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.9753 -0.5331 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.9753 -0.5331 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.6898 -0.9456 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.6898 -0.9456 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.4043 -0.5331 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.4043 -0.5331 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.4043 0.2918 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.4043 0.2918 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.6898 0.7043 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.6898 0.7043 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.9753 0.2918 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.9753 0.2918 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.3372 -0.4716 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.3372 -0.4716 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1714 -2.9599 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1714 -2.9599 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.0836 0.6841 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.0836 0.6841 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.6898 1.5293 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.6898 1.5293 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.5237 2.1285 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.5237 2.1285 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.0521 1.6002 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.0521 1.6002 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6820 0.9509 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6820 0.9509 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6820 0.2038 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6820 0.2038 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1536 0.7321 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1536 0.7321 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.5237 1.3814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.5237 1.3814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.8658 2.7210 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.8658 2.7210 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.0714 2.3568 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.0714 2.3568 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1551 0.2137 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1551 0.2137 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.9023 1.3922 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.9023 1.3922 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.9335 1.3922 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.9335 1.3922 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.3236 0.9531 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.3236 0.9531 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.4484 0.4370 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.4484 0.4370 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.5285 0.9082 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.5285 0.9082 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.9023 1.9082 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.9023 1.9082 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.9335 1.9078 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.9335 1.9078 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.4285 2.2997 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.4285 2.2997 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.0836 1.9215 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.0836 1.9215 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.4285 2.9803 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.4285 2.9803 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.9335 1.0003 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.9335 1.0003 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.8311 1.0003 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.8311 1.0003 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 18 1 1 0 0 0 0 | + | 18 1 1 0 0 0 0 |
| − | 3 19 1 0 0 0 0 | + | 3 19 1 0 0 0 0 |
| − | 15 20 1 0 0 0 0 | + | 15 20 1 0 0 0 0 |
| − | 16 21 1 0 0 0 0 | + | 16 21 1 0 0 0 0 |
| − | 22 23 1 1 0 0 0 | + | 22 23 1 1 0 0 0 |
| − | 23 24 1 1 0 0 0 | + | 23 24 1 1 0 0 0 |
| − | 25 24 1 1 0 0 0 | + | 25 24 1 1 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 27 1 0 0 0 0 | + | 26 27 1 0 0 0 0 |
| − | 27 22 1 0 0 0 0 | + | 27 22 1 0 0 0 0 |
| − | 22 28 1 0 0 0 0 | + | 22 28 1 0 0 0 0 |
| − | 23 29 1 0 0 0 0 | + | 23 29 1 0 0 0 0 |
| − | 25 18 1 0 0 0 0 | + | 25 18 1 0 0 0 0 |
| − | 24 30 1 0 0 0 0 | + | 24 30 1 0 0 0 0 |
| − | 31 32 1 0 0 0 0 | + | 31 32 1 0 0 0 0 |
| − | 32 33 1 0 0 0 0 | + | 32 33 1 0 0 0 0 |
| − | 33 34 1 1 0 0 0 | + | 33 34 1 1 0 0 0 |
| − | 35 34 1 1 0 0 0 | + | 35 34 1 1 0 0 0 |
| − | 35 31 1 0 0 0 0 | + | 35 31 1 0 0 0 0 |
| − | 31 36 1 0 0 0 0 | + | 31 36 1 0 0 0 0 |
| − | 35 30 1 0 0 0 0 | + | 35 30 1 0 0 0 0 |
| − | 32 37 1 0 0 0 0 | + | 32 37 1 0 0 0 0 |
| − | 37 38 1 0 0 0 0 | + | 37 38 1 0 0 0 0 |
| − | 38 39 2 0 0 0 0 | + | 38 39 2 0 0 0 0 |
| − | 38 40 1 0 0 0 0 | + | 38 40 1 0 0 0 0 |
| − | 41 42 1 0 0 0 0 | + | 41 42 1 0 0 0 0 |
| − | 32 41 1 0 0 0 0 | + | 32 41 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 41 42 | + | M SAL 1 2 41 42 |
| − | M SBL 1 1 46 | + | M SBL 1 1 46 |
| − | M SMT 1 CH2OH | + | M SMT 1 CH2OH |
| − | M SBV 1 46 0.0000 0.3919 | + | M SBV 1 46 0.0000 0.3919 |
| − | S SKP 5 | + | S SKP 5 |
| − | ID FL3FACGS0056 | + | ID FL3FACGS0056 |
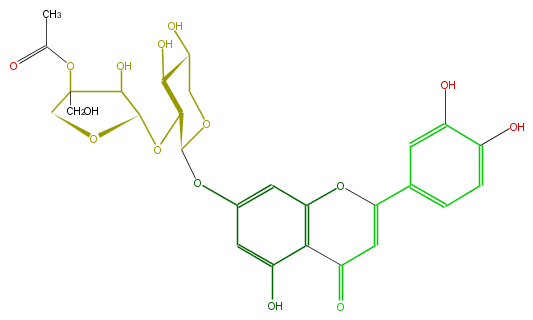
| − | FORMULA C27H28O15 | + | FORMULA C27H28O15 |
| − | EXACTMASS 592.1428202259999 | + | EXACTMASS 592.1428202259999 |
| − | AVERAGEMASS 592.5022200000001 | + | AVERAGEMASS 592.5022200000001 |
| − | SMILES c(c1)c(c(O)cc1C(=C5)Oc(c4C(=O)5)cc(cc(O)4)OC(C(OC(O3)C(C(CO)(OC(C)=O)C3)O)2)OCC(O)C(O)2)O | + | SMILES c(c1)c(c(O)cc1C(=C5)Oc(c4C(=O)5)cc(cc(O)4)OC(C(OC(O3)C(C(CO)(OC(C)=O)C3)O)2)OCC(O)C(O)2)O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
42 46 0 0 0 0 0 0 0 0999 V2000
-0.5298 -0.9377 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5298 -1.7473 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.1714 -2.1520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.8723 -1.7473 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.8723 -0.9377 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.1714 -0.5330 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.5734 -2.1520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.2745 -1.7473 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.2745 -0.9377 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.5734 -0.5330 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.5734 -2.9803 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.9753 -0.5331 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.6898 -0.9456 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.4043 -0.5331 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.4043 0.2918 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.6898 0.7043 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.9753 0.2918 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.3372 -0.4716 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.1714 -2.9599 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.0836 0.6841 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.6898 1.5293 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.5237 2.1285 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.0521 1.6002 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6820 0.9509 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6820 0.2038 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1536 0.7321 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.5237 1.3814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.8658 2.7210 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.0714 2.3568 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.1551 0.2137 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.9023 1.3922 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.9335 1.3922 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.3236 0.9531 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.4484 0.4370 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.5285 0.9082 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.9023 1.9082 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.9335 1.9078 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.4285 2.2997 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.0836 1.9215 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.4285 2.9803 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.9335 1.0003 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.8311 1.0003 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
18 1 1 0 0 0 0
3 19 1 0 0 0 0
15 20 1 0 0 0 0
16 21 1 0 0 0 0
22 23 1 1 0 0 0
23 24 1 1 0 0 0
25 24 1 1 0 0 0
25 26 1 0 0 0 0
26 27 1 0 0 0 0
27 22 1 0 0 0 0
22 28 1 0 0 0 0
23 29 1 0 0 0 0
25 18 1 0 0 0 0
24 30 1 0 0 0 0
31 32 1 0 0 0 0
32 33 1 0 0 0 0
33 34 1 1 0 0 0
35 34 1 1 0 0 0
35 31 1 0 0 0 0
31 36 1 0 0 0 0
35 30 1 0 0 0 0
32 37 1 0 0 0 0
37 38 1 0 0 0 0
38 39 2 0 0 0 0
38 40 1 0 0 0 0
41 42 1 0 0 0 0
32 41 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 41 42
M SBL 1 1 46
M SMT 1 CH2OH
M SBV 1 46 0.0000 0.3919
S SKP 5
ID FL3FACGS0056
FORMULA C27H28O15
EXACTMASS 592.1428202259999
AVERAGEMASS 592.5022200000001
SMILES c(c1)c(c(O)cc1C(=C5)Oc(c4C(=O)5)cc(cc(O)4)OC(C(OC(O3)C(C(CO)(OC(C)=O)C3)O)2)OCC(O)C(O)2)O
M END