Mol:FL3FACGS0045
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 53 58 0 0 0 0 0 0 0 0999 V2000 | + | 53 58 0 0 0 0 0 0 0 0999 V2000 |
| − | -0.4089 -1.0942 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.4089 -1.0942 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.4089 -1.7577 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.4089 -1.7577 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1659 -2.0895 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1659 -2.0895 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7404 -1.7577 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7404 -1.7577 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7404 -1.0942 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7404 -1.0942 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1659 -0.7625 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1659 -0.7625 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3150 -2.0895 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3150 -2.0895 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8895 -1.7577 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8895 -1.7577 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8895 -1.0942 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8895 -1.0942 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3150 -0.7625 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3150 -0.7625 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5337 -2.5582 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5337 -2.5582 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.4639 -0.7626 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.4639 -0.7626 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0496 -1.1007 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0496 -1.1007 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.6352 -0.7626 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.6352 -0.7626 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.6352 -0.0863 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.6352 -0.0863 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0496 0.2518 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0496 0.2518 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.4639 -0.0863 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.4639 -0.0863 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0706 -0.7121 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0706 -0.7121 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1659 -2.7515 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1659 -2.7515 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.3923 0.3508 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.3923 0.3508 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.2207 -1.1007 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.2207 -1.1007 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.8456 1.6331 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.8456 1.6331 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.6460 1.1711 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.6460 1.1711 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.3921 2.0596 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.3921 2.0596 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.6460 2.9535 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.6460 2.9535 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.8456 3.4157 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.8456 3.4157 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.0995 2.5272 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.0995 2.5272 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.1504 3.9436 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.1504 3.9436 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.6460 4.0482 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.6460 4.0482 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.8985 1.5532 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.8985 1.5532 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.2909 -0.5218 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.2909 -0.5218 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.6339 -1.3890 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.6339 -1.3890 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.6881 -1.0211 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.6881 -1.0211 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.7755 -1.0112 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.7755 -1.0112 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.4387 -0.3479 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.4387 -0.3479 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.4050 -0.6948 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.4050 -0.6948 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.8985 -0.8726 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.8985 -0.8726 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.3159 -1.3890 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.3159 -1.3890 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.7965 -1.4938 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.7965 -1.4938 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.4714 -2.3596 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.4714 -2.3596 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.9652 -3.2149 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.9652 -3.2149 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.0155 -2.9435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.0155 -2.9435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.0602 -3.2149 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.0602 -3.2149 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.5662 -2.3596 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.5662 -2.3596 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.5161 -2.6309 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.5161 -2.6309 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.3422 -2.3596 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.3422 -2.3596 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.4155 -2.9549 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.4155 -2.9549 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.5718 -3.4999 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.5718 -3.4999 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.0602 -4.0482 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.0602 -4.0482 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.9883 -0.1114 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.9883 -0.1114 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.0799 0.9356 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.0799 0.9356 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.4765 3.1502 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.4765 3.1502 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.5054 2.3258 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5054 2.3258 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 18 1 1 0 0 0 0 | + | 18 1 1 0 0 0 0 |
| − | 3 19 1 0 0 0 0 | + | 3 19 1 0 0 0 0 |
| − | 15 20 1 0 0 0 0 | + | 15 20 1 0 0 0 0 |
| − | 14 21 1 0 0 0 0 | + | 14 21 1 0 0 0 0 |
| − | 23 22 1 1 0 0 0 | + | 23 22 1 1 0 0 0 |
| − | 23 24 1 0 0 0 0 | + | 23 24 1 0 0 0 0 |
| − | 24 25 1 0 0 0 0 | + | 24 25 1 0 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 27 1 1 0 0 0 | + | 26 27 1 1 0 0 0 |
| − | 27 22 1 1 0 0 0 | + | 27 22 1 1 0 0 0 |
| − | 26 28 1 0 0 0 0 | + | 26 28 1 0 0 0 0 |
| − | 25 29 1 0 0 0 0 | + | 25 29 1 0 0 0 0 |
| − | 24 30 1 0 0 0 0 | + | 24 30 1 0 0 0 0 |
| − | 20 23 1 0 0 0 0 | + | 20 23 1 0 0 0 0 |
| − | 31 32 1 1 0 0 0 | + | 31 32 1 1 0 0 0 |
| − | 32 33 1 1 0 0 0 | + | 32 33 1 1 0 0 0 |
| − | 34 33 1 1 0 0 0 | + | 34 33 1 1 0 0 0 |
| − | 34 35 1 0 0 0 0 | + | 34 35 1 0 0 0 0 |
| − | 35 36 1 0 0 0 0 | + | 35 36 1 0 0 0 0 |
| − | 36 31 1 0 0 0 0 | + | 36 31 1 0 0 0 0 |
| − | 31 37 1 0 0 0 0 | + | 31 37 1 0 0 0 0 |
| − | 32 38 1 0 0 0 0 | + | 32 38 1 0 0 0 0 |
| − | 33 39 1 0 0 0 0 | + | 33 39 1 0 0 0 0 |
| − | 40 41 1 0 0 0 0 | + | 40 41 1 0 0 0 0 |
| − | 41 42 1 1 0 0 0 | + | 41 42 1 1 0 0 0 |
| − | 42 43 1 1 0 0 0 | + | 42 43 1 1 0 0 0 |
| − | 44 43 1 1 0 0 0 | + | 44 43 1 1 0 0 0 |
| − | 44 45 1 0 0 0 0 | + | 44 45 1 0 0 0 0 |
| − | 45 40 1 0 0 0 0 | + | 45 40 1 0 0 0 0 |
| − | 40 46 1 0 0 0 0 | + | 40 46 1 0 0 0 0 |
| − | 41 47 1 0 0 0 0 | + | 41 47 1 0 0 0 0 |
| − | 42 48 1 0 0 0 0 | + | 42 48 1 0 0 0 0 |
| − | 43 49 1 0 0 0 0 | + | 43 49 1 0 0 0 0 |
| − | 44 39 1 0 0 0 0 | + | 44 39 1 0 0 0 0 |
| − | 34 18 1 0 0 0 0 | + | 34 18 1 0 0 0 0 |
| − | 50 51 1 0 0 0 0 | + | 50 51 1 0 0 0 0 |
| − | 36 50 1 0 0 0 0 | + | 36 50 1 0 0 0 0 |
| − | 52 53 1 0 0 0 0 | + | 52 53 1 0 0 0 0 |
| − | 27 52 1 0 0 0 0 | + | 27 52 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 50 51 | + | M SAL 1 2 50 51 |
| − | M SBL 1 1 56 | + | M SBL 1 1 56 |
| − | M SMT 1 ^ CH2OH | + | M SMT 1 ^ CH2OH |
| − | M SBV 1 56 0.5833 -0.5834 | + | M SBV 1 56 0.5833 -0.5834 |
| − | M STY 1 2 SUP | + | M STY 1 2 SUP |
| − | M SLB 1 2 2 | + | M SLB 1 2 2 |
| − | M SAL 2 2 52 53 | + | M SAL 2 2 52 53 |
| − | M SBL 2 1 58 | + | M SBL 2 1 58 |
| − | M SMT 2 ^ CH2OH | + | M SMT 2 ^ CH2OH |
| − | M SBV 2 58 0.6230 -0.6230 | + | M SBV 2 58 0.6230 -0.6230 |
| − | S SKP 5 | + | S SKP 5 |
| − | ID FL3FACGS0045 | + | ID FL3FACGS0045 |
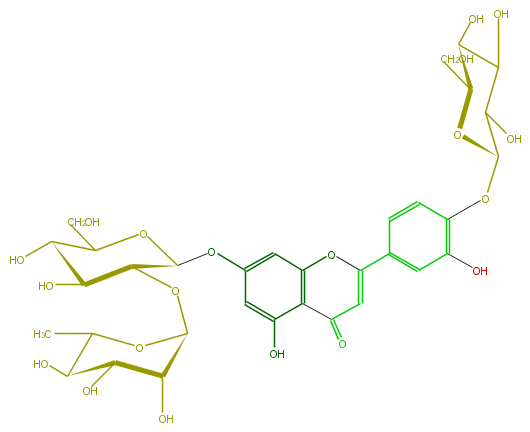
| − | FORMULA C33H40O20 | + | FORMULA C33H40O20 |
| − | EXACTMASS 756.21129372 | + | EXACTMASS 756.21129372 |
| − | AVERAGEMASS 756.6587 | + | AVERAGEMASS 756.6587 |
| − | SMILES OC(C(OC(O6)C(C(C(O)C6C)O)O)1)C(C(OC(Oc(c5)cc(c(c52)C(C=C(c(c4)cc(c(c4)OC(O3)C(C(C(C3CO)O)O)O)O)O2)=O)O)1)CO)O | + | SMILES OC(C(OC(O6)C(C(C(O)C6C)O)O)1)C(C(OC(Oc(c5)cc(c(c52)C(C=C(c(c4)cc(c(c4)OC(O3)C(C(C(C3CO)O)O)O)O)O2)=O)O)1)CO)O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
53 58 0 0 0 0 0 0 0 0999 V2000
-0.4089 -1.0942 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.4089 -1.7577 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.1659 -2.0895 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7404 -1.7577 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7404 -1.0942 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.1659 -0.7625 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.3150 -2.0895 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8895 -1.7577 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8895 -1.0942 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.3150 -0.7625 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.5337 -2.5582 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.4639 -0.7626 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.0496 -1.1007 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.6352 -0.7626 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.6352 -0.0863 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.0496 0.2518 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.4639 -0.0863 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.0706 -0.7121 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.1659 -2.7515 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.3923 0.3508 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.2207 -1.1007 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.8456 1.6331 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.6460 1.1711 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3921 2.0596 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.6460 2.9535 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.8456 3.4157 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.0995 2.5272 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.1504 3.9436 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.6460 4.0482 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.8985 1.5532 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.2909 -0.5218 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.6339 -1.3890 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.6881 -1.0211 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.7755 -1.0112 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.4387 -0.3479 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.4050 -0.6948 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.8985 -0.8726 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.3159 -1.3890 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.7965 -1.4938 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.4714 -2.3596 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.9652 -3.2149 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.0155 -2.9435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.0602 -3.2149 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.5662 -2.3596 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.5161 -2.6309 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.3422 -2.3596 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.4155 -2.9549 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.5718 -3.4999 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.0602 -4.0482 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.9883 -0.1114 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.0799 0.9356 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.4765 3.1502 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.5054 2.3258 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
18 1 1 0 0 0 0
3 19 1 0 0 0 0
15 20 1 0 0 0 0
14 21 1 0 0 0 0
23 22 1 1 0 0 0
23 24 1 0 0 0 0
24 25 1 0 0 0 0
25 26 1 0 0 0 0
26 27 1 1 0 0 0
27 22 1 1 0 0 0
26 28 1 0 0 0 0
25 29 1 0 0 0 0
24 30 1 0 0 0 0
20 23 1 0 0 0 0
31 32 1 1 0 0 0
32 33 1 1 0 0 0
34 33 1 1 0 0 0
34 35 1 0 0 0 0
35 36 1 0 0 0 0
36 31 1 0 0 0 0
31 37 1 0 0 0 0
32 38 1 0 0 0 0
33 39 1 0 0 0 0
40 41 1 0 0 0 0
41 42 1 1 0 0 0
42 43 1 1 0 0 0
44 43 1 1 0 0 0
44 45 1 0 0 0 0
45 40 1 0 0 0 0
40 46 1 0 0 0 0
41 47 1 0 0 0 0
42 48 1 0 0 0 0
43 49 1 0 0 0 0
44 39 1 0 0 0 0
34 18 1 0 0 0 0
50 51 1 0 0 0 0
36 50 1 0 0 0 0
52 53 1 0 0 0 0
27 52 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 50 51
M SBL 1 1 56
M SMT 1 ^ CH2OH
M SBV 1 56 0.5833 -0.5834
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 52 53
M SBL 2 1 58
M SMT 2 ^ CH2OH
M SBV 2 58 0.6230 -0.6230
S SKP 5
ID FL3FACGS0045
FORMULA C33H40O20
EXACTMASS 756.21129372
AVERAGEMASS 756.6587
SMILES OC(C(OC(O6)C(C(C(O)C6C)O)O)1)C(C(OC(Oc(c5)cc(c(c52)C(C=C(c(c4)cc(c(c4)OC(O3)C(C(C(C3CO)O)O)O)O)O2)=O)O)1)CO)O
M END