Mol:PR100346
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | ACD/Labs08070914182D | + | ACD/Labs08070914182D |
| − | + | ||
| − | 46 50 0 0 1 0 0 0 0 0 2 V2000 | + | 46 50 0 0 1 0 0 0 0 0 2 V2000 |
| − | 20.6257 -6.5167 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 20.6257 -6.5167 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 20.6188 -7.3477 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 20.6188 -7.3477 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 21.3395 -6.1098 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 21.3395 -6.1098 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 19.9084 -6.0995 0.0000 O 0 3 0 0 0 0 0 0 0 0 0 0 | + | 19.9084 -6.0995 0.0000 O 0 3 0 0 0 0 0 0 0 0 0 0 |
| − | 19.9015 -7.7546 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 19.9015 -7.7546 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 21.3326 -7.7615 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 21.3326 -7.7615 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.0498 -6.5270 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.0498 -6.5270 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 21.3429 -5.2857 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 21.3429 -5.2857 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 19.1877 -6.5064 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 19.1877 -6.5064 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 19.1843 -7.3374 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 19.1843 -7.3374 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.1636 -8.2995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.1636 -8.2995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.7671 -6.1167 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.7671 -6.1167 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.0567 -4.8753 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.0567 -4.8753 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 18.4705 -6.0960 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 18.4705 -6.0960 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 18.4705 -7.7546 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 18.4705 -7.7546 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.1567 -9.1202 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.1567 -9.1202 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.8808 -7.8891 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.8808 -7.8891 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.7705 -5.2926 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.7705 -5.2926 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 23.4774 -6.5339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 23.4774 -6.5339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.0636 -4.0512 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.0636 -4.0512 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 17.7498 -6.5064 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 17.7498 -6.5064 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 18.4705 -8.5615 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 18.4705 -8.5615 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 17.7498 -7.3408 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 17.7498 -7.3408 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.8705 -9.5374 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.8705 -9.5374 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 23.5912 -8.3064 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 23.5912 -8.3064 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.8843 -7.0650 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.8843 -7.0650 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 23.4877 -4.8857 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 23.4877 -4.8857 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 24.1946 -6.1270 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 24.1946 -6.1270 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 20.8049 -3.4291 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 20.8049 -3.4291 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 17.0360 -6.0926 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 17.0360 -6.0926 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 17.6808 -9.0581 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 17.6808 -9.0581 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 23.5843 -9.1339 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 23.5843 -9.1339 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.8636 -10.3650 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.8636 -10.3650 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 24.3084 -7.8995 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 24.3084 -7.8995 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 16.9705 -8.6443 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 16.9705 -8.6443 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 17.6808 -9.8822 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 17.6808 -9.8822 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 24.2981 -9.5512 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 24.2981 -9.5512 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.1464 -10.7719 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.1464 -10.7719 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 16.2533 -9.0581 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 16.2533 -9.0581 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 16.9705 -10.2960 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 16.9705 -10.2960 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 18.3981 -10.2960 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 18.3981 -10.2960 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 16.2533 -9.8822 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 16.2533 -9.8822 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 15.5395 -8.6443 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 15.5395 -8.6443 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 16.9705 -11.1202 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 16.9705 -11.1202 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 15.4434 -10.4883 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 15.4434 -10.4883 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 14.8257 -9.0581 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 14.8257 -9.0581 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 1 3 1 0 0 0 0 | + | 1 3 1 0 0 0 0 |
| − | 1 4 2 0 0 0 0 | + | 1 4 2 0 0 0 0 |
| − | 2 5 2 0 0 0 0 | + | 2 5 2 0 0 0 0 |
| − | 2 6 1 0 0 0 0 | + | 2 6 1 0 0 0 0 |
| − | 3 7 2 0 0 0 0 | + | 3 7 2 0 0 0 0 |
| − | 3 8 1 0 0 0 0 | + | 3 8 1 0 0 0 0 |
| − | 4 9 1 0 0 0 0 | + | 4 9 1 0 0 0 0 |
| − | 5 10 1 0 0 0 0 | + | 5 10 1 0 0 0 0 |
| − | 11 6 1 1 0 0 0 | + | 11 6 1 1 0 0 0 |
| − | 7 12 1 0 0 0 0 | + | 7 12 1 0 0 0 0 |
| − | 8 13 2 0 0 0 0 | + | 8 13 2 0 0 0 0 |
| − | 9 14 1 0 0 0 0 | + | 9 14 1 0 0 0 0 |
| − | 10 15 1 0 0 0 0 | + | 10 15 1 0 0 0 0 |
| − | 11 16 1 0 0 0 0 | + | 11 16 1 0 0 0 0 |
| − | 11 17 1 0 0 0 0 | + | 11 17 1 0 0 0 0 |
| − | 12 18 2 0 0 0 0 | + | 12 18 2 0 0 0 0 |
| − | 12 19 1 0 0 0 0 | + | 12 19 1 0 0 0 0 |
| − | 13 20 1 0 0 0 0 | + | 13 20 1 0 0 0 0 |
| − | 14 21 2 0 0 0 0 | + | 14 21 2 0 0 0 0 |
| − | 15 22 1 0 0 0 0 | + | 15 22 1 0 0 0 0 |
| − | 15 23 2 0 0 0 0 | + | 15 23 2 0 0 0 0 |
| − | 16 24 1 0 0 0 0 | + | 16 24 1 0 0 0 0 |
| − | 17 25 1 0 0 0 0 | + | 17 25 1 0 0 0 0 |
| − | 17 26 1 6 0 0 0 | + | 17 26 1 6 0 0 0 |
| − | 18 27 1 0 0 0 0 | + | 18 27 1 0 0 0 0 |
| − | 19 28 1 0 0 0 0 | + | 19 28 1 0 0 0 0 |
| − | 20 29 1 0 0 0 0 | + | 20 29 1 0 0 0 0 |
| − | 21 30 1 0 0 0 0 | + | 21 30 1 0 0 0 0 |
| − | 31 22 1 1 0 0 0 | + | 31 22 1 1 0 0 0 |
| − | 24 32 1 0 0 0 0 | + | 24 32 1 0 0 0 0 |
| − | 24 33 1 1 0 0 0 | + | 24 33 1 1 0 0 0 |
| − | 25 34 1 1 0 0 0 | + | 25 34 1 1 0 0 0 |
| − | 31 35 1 0 0 0 0 | + | 31 35 1 0 0 0 0 |
| − | 31 36 1 0 0 0 0 | + | 31 36 1 0 0 0 0 |
| − | 32 37 1 6 0 0 0 | + | 32 37 1 6 0 0 0 |
| − | 33 38 1 0 0 0 0 | + | 33 38 1 0 0 0 0 |
| − | 35 39 1 0 0 0 0 | + | 35 39 1 0 0 0 0 |
| − | 36 40 1 0 0 0 0 | + | 36 40 1 0 0 0 0 |
| − | 36 41 1 6 0 0 0 | + | 36 41 1 6 0 0 0 |
| − | 39 42 1 0 0 0 0 | + | 39 42 1 0 0 0 0 |
| − | 39 43 1 1 0 0 0 | + | 39 43 1 1 0 0 0 |
| − | 40 44 1 1 0 0 0 | + | 40 44 1 1 0 0 0 |
| − | 42 45 1 6 0 0 0 | + | 42 45 1 6 0 0 0 |
| − | 43 46 1 0 0 0 0 | + | 43 46 1 0 0 0 0 |
| − | 9 10 2 0 0 0 0 | + | 9 10 2 0 0 0 0 |
| − | 13 18 1 0 0 0 0 | + | 13 18 1 0 0 0 0 |
| − | 21 23 1 0 0 0 0 | + | 21 23 1 0 0 0 0 |
| − | 25 32 1 0 0 0 0 | + | 25 32 1 0 0 0 0 |
| − | 40 42 1 0 0 0 0 | + | 40 42 1 0 0 0 0 |
| − | M CHG 1 4 1 | + | M CHG 1 4 1 |
| − | S SKP | + | S SKP 7 |
| − | CAS_RN 47863-30-9 | + | CAS_RN 47863-30-9 |
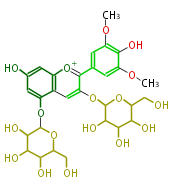
| − | NAME Malvin | + | NAME Malvin |
| + | ID PR100346 | ||
| + | FORMULA C29H35O17 | ||
| + | EXACTMASS 655.187424694 | ||
| + | AVERAGEMASS 655.578 | ||
| + | SMILES c(c41)([o+1]c(c(OC(O5)C(O)C(O)C(O)C5CO)c4)c(c3)cc(OC)c(O)c(OC)3)cc(O)cc1OC(O2)C(O)C(O)C(O)C2CO | ||
M END | M END | ||
Latest revision as of 16:44, 14 January 2011
ACD/Labs08070914182D 46 50 0 0 1 0 0 0 0 0 2 V2000 20.6257 -6.5167 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 20.6188 -7.3477 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 21.3395 -6.1098 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 19.9084 -6.0995 0.0000 O 0 3 0 0 0 0 0 0 0 0 0 0 19.9015 -7.7546 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 21.3326 -7.7615 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 22.0498 -6.5270 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 21.3429 -5.2857 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 19.1877 -6.5064 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 19.1843 -7.3374 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 22.1636 -8.2995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 22.7671 -6.1167 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 22.0567 -4.8753 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 18.4705 -6.0960 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 18.4705 -7.7546 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 22.1567 -9.1202 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 22.8808 -7.8891 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 22.7705 -5.2926 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 23.4774 -6.5339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 22.0636 -4.0512 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 17.7498 -6.5064 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 18.4705 -8.5615 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 17.7498 -7.3408 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 22.8705 -9.5374 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 23.5912 -8.3064 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 22.8843 -7.0650 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 23.4877 -4.8857 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 24.1946 -6.1270 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 20.8049 -3.4291 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 17.0360 -6.0926 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 17.6808 -9.0581 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 23.5843 -9.1339 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 22.8636 -10.3650 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 24.3084 -7.8995 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 16.9705 -8.6443 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 17.6808 -9.8822 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 24.2981 -9.5512 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 22.1464 -10.7719 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 16.2533 -9.0581 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 16.9705 -10.2960 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 18.3981 -10.2960 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 16.2533 -9.8822 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 15.5395 -8.6443 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 16.9705 -11.1202 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 15.4434 -10.4883 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 14.8257 -9.0581 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 1 2 1 0 0 0 0 1 3 1 0 0 0 0 1 4 2 0 0 0 0 2 5 2 0 0 0 0 2 6 1 0 0 0 0 3 7 2 0 0 0 0 3 8 1 0 0 0 0 4 9 1 0 0 0 0 5 10 1 0 0 0 0 11 6 1 1 0 0 0 7 12 1 0 0 0 0 8 13 2 0 0 0 0 9 14 1 0 0 0 0 10 15 1 0 0 0 0 11 16 1 0 0 0 0 11 17 1 0 0 0 0 12 18 2 0 0 0 0 12 19 1 0 0 0 0 13 20 1 0 0 0 0 14 21 2 0 0 0 0 15 22 1 0 0 0 0 15 23 2 0 0 0 0 16 24 1 0 0 0 0 17 25 1 0 0 0 0 17 26 1 6 0 0 0 18 27 1 0 0 0 0 19 28 1 0 0 0 0 20 29 1 0 0 0 0 21 30 1 0 0 0 0 31 22 1 1 0 0 0 24 32 1 0 0 0 0 24 33 1 1 0 0 0 25 34 1 1 0 0 0 31 35 1 0 0 0 0 31 36 1 0 0 0 0 32 37 1 6 0 0 0 33 38 1 0 0 0 0 35 39 1 0 0 0 0 36 40 1 0 0 0 0 36 41 1 6 0 0 0 39 42 1 0 0 0 0 39 43 1 1 0 0 0 40 44 1 1 0 0 0 42 45 1 6 0 0 0 43 46 1 0 0 0 0 9 10 2 0 0 0 0 13 18 1 0 0 0 0 21 23 1 0 0 0 0 25 32 1 0 0 0 0 40 42 1 0 0 0 0 M CHG 1 4 1 S SKP 7 CAS_RN 47863-30-9 NAME Malvin ID PR100346 FORMULA C29H35O17 EXACTMASS 655.187424694 AVERAGEMASS 655.578 SMILES c(c41)([o+1]c(c(OC(O5)C(O)C(O)C(O)C5CO)c4)c(c3)cc(OC)c(O)c(OC)3)cc(O)cc1OC(O2)C(O)C(O)C(O)C2CO M END