Mol:BMMCBZ1Sa038
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 58 61 0 0 1 0 0 0 0 0999 V2000 | + | 58 61 0 0 1 0 0 0 0 0999 V2000 |
| − | 6.5817 -3.5672 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 6.5817 -3.5672 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.6036 -3.3593 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.6036 -3.3593 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.9563 -3.8945 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.9563 -3.8945 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.6473 -4.8456 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.6473 -4.8456 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.6691 -5.0535 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.6691 -5.0535 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0000 -4.3103 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0000 -4.3103 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.3090 -3.3593 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.3090 -3.3593 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.2872 -3.1514 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.2872 -3.1514 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.9344 -4.1024 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.9344 -4.1024 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 6.8907 -4.5183 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 6.8907 -4.5183 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 23.9388 -2.5373 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 | + | 23.9388 -2.5373 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 23.0728 -2.0373 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 23.0728 -2.0373 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 23.0728 -1.0373 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 | + | 23.0728 -1.0373 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 23.9388 -0.5373 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 23.9388 -0.5373 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 24.8049 -1.0373 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 24.8049 -1.0373 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 24.8049 -2.0373 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 24.8049 -2.0373 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 25.5480 -0.3681 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 | + | 25.5480 -0.3681 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 25.1413 0.5454 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 25.1413 0.5454 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 24.1467 0.4409 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 | + | 24.1467 0.4409 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 25.6709 -2.5373 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 | + | 25.6709 -2.5373 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 23.4776 1.1840 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 23.4776 1.1840 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 23.6855 2.1622 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 23.6855 2.1622 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 22.8195 2.6622 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 22.8195 2.6622 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 22.0764 1.9930 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 22.0764 1.9930 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 21.0982 2.2010 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 21.0982 2.2010 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 24.5991 2.5689 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 24.5991 2.5689 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.7150 3.6567 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.7150 3.6567 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.4831 1.0795 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.4831 1.0795 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 20.4291 1.4578 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 20.4291 1.4578 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 23.5240 4.2445 0.0000 P 0 0 0 0 0 0 0 0 0 0 0 0 | + | 23.5240 4.2445 0.0000 P 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 24.1118 3.4355 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 24.1118 3.4355 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.9362 5.0535 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.9362 5.0535 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 24.3330 4.8323 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 24.3330 4.8323 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 19.4509 1.6657 0.0000 P 0 0 0 0 0 0 0 0 0 0 0 0 | + | 19.4509 1.6657 0.0000 P 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 19.2430 0.6876 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 19.2430 0.6876 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 19.6588 2.6439 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 19.6588 2.6439 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 18.4728 1.8736 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 18.4728 1.8736 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 17.8037 1.1305 0.0000 P 0 0 0 0 0 0 0 0 0 0 0 0 | + | 17.8037 1.1305 0.0000 P 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 18.5468 0.4614 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 18.5468 0.4614 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 17.0605 1.7996 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 17.0605 1.7996 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 17.1345 0.3873 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 17.1345 0.3873 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 16.1564 0.5953 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 16.1564 0.5953 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 15.4872 -0.1479 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 15.4872 -0.1479 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 16.2304 -0.8170 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 16.2304 -0.8170 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 14.7441 0.5212 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 14.7441 0.5212 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 14.8181 -0.8910 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 | + | 14.8181 -0.8910 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 |
| − | 13.8400 -0.6831 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 13.8400 -0.6831 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 13.1708 -1.4263 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 | + | 13.1708 -1.4263 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 12.1927 -1.2184 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 12.1927 -1.2184 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 11.5236 -1.9615 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 11.5236 -1.9615 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 10.5454 -1.7536 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 10.5454 -1.7536 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 9.8763 -2.4967 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 | + | 9.8763 -2.4967 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 8.8981 -2.2888 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 8.8981 -2.2888 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 8.2290 -3.0320 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 8.2290 -3.0320 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 7.2509 -2.8241 0.0000 S 0 0 0 0 0 0 0 0 0 0 0 0 | + | 7.2509 -2.8241 0.0000 S 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 15.1271 -1.8421 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 15.1271 -1.8421 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 13.5309 0.2679 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 13.5309 0.2679 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 10.2364 -0.8025 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 10.2364 -0.8025 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 11 16 2 0 0 0 0 | + | 11 16 2 0 0 0 0 |
| − | 11 12 1 0 0 0 0 | + | 11 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 15 14 2 0 0 0 0 | + | 15 14 2 0 0 0 0 |
| − | 19 18 1 0 0 0 0 | + | 19 18 1 0 0 0 0 |
| − | 18 17 2 0 0 0 0 | + | 18 17 2 0 0 0 0 |
| − | 17 15 1 0 0 0 0 | + | 17 15 1 0 0 0 0 |
| − | 14 19 1 0 0 0 0 | + | 14 19 1 0 0 0 0 |
| − | 16 20 1 0 0 0 0 | + | 16 20 1 0 0 0 0 |
| − | 24 28 1 6 0 0 0 | + | 24 28 1 6 0 0 0 |
| − | 23 24 1 0 0 0 0 | + | 23 24 1 0 0 0 0 |
| − | 21 28 1 6 0 0 0 | + | 21 28 1 6 0 0 0 |
| − | 23 22 1 0 0 0 0 | + | 23 22 1 0 0 0 0 |
| − | 21 22 1 0 0 0 0 | + | 21 22 1 0 0 0 0 |
| − | 24 25 1 0 0 0 0 | + | 24 25 1 0 0 0 0 |
| − | 25 29 1 0 0 0 0 | + | 25 29 1 0 0 0 0 |
| − | 21 19 1 0 0 0 0 | + | 21 19 1 0 0 0 0 |
| − | 29 34 1 0 0 0 0 | + | 29 34 1 0 0 0 0 |
| − | 34 36 1 0 0 0 0 | + | 34 36 1 0 0 0 0 |
| − | 34 35 2 0 0 0 0 | + | 34 35 2 0 0 0 0 |
| − | 34 37 1 0 0 0 0 | + | 34 37 1 0 0 0 0 |
| − | 41 38 1 0 0 0 0 | + | 41 38 1 0 0 0 0 |
| − | 38 40 2 0 0 0 0 | + | 38 40 2 0 0 0 0 |
| − | 38 39 1 0 0 0 0 | + | 38 39 1 0 0 0 0 |
| − | 41 42 1 0 0 0 0 | + | 41 42 1 0 0 0 0 |
| − | 38 37 1 0 0 0 0 | + | 38 37 1 0 0 0 0 |
| − | 42 43 1 0 0 0 0 | + | 42 43 1 0 0 0 0 |
| − | 43 46 1 0 0 0 0 | + | 43 46 1 0 0 0 0 |
| − | 43 45 1 0 0 0 0 | + | 43 45 1 0 0 0 0 |
| − | 43 44 1 0 0 0 0 | + | 43 44 1 0 0 0 0 |
| − | 46 47 1 0 0 0 0 | + | 46 47 1 0 0 0 0 |
| − | 46 56 1 1 0 0 0 | + | 46 56 1 1 0 0 0 |
| − | 47 57 2 0 0 0 0 | + | 47 57 2 0 0 0 0 |
| − | 47 48 1 0 0 0 0 | + | 47 48 1 0 0 0 0 |
| − | 48 49 1 0 0 0 0 | + | 48 49 1 0 0 0 0 |
| − | 49 50 1 0 0 0 0 | + | 49 50 1 0 0 0 0 |
| − | 50 51 1 0 0 0 0 | + | 50 51 1 0 0 0 0 |
| − | 51 52 1 0 0 0 0 | + | 51 52 1 0 0 0 0 |
| − | 51 58 2 0 0 0 0 | + | 51 58 2 0 0 0 0 |
| − | 52 53 1 0 0 0 0 | + | 52 53 1 0 0 0 0 |
| − | 53 54 1 0 0 0 0 | + | 53 54 1 0 0 0 0 |
| − | 22 26 1 1 0 0 0 | + | 22 26 1 1 0 0 0 |
| − | 23 27 1 1 0 0 0 | + | 23 27 1 1 0 0 0 |
| − | 31 30 1 0 0 0 0 | + | 31 30 1 0 0 0 0 |
| − | 30 33 1 0 0 0 0 | + | 30 33 1 0 0 0 0 |
| − | 30 32 2 0 0 0 0 | + | 30 32 2 0 0 0 0 |
| − | 27 30 1 0 0 0 0 | + | 27 30 1 0 0 0 0 |
| − | 8 3 2 0 0 0 0 | + | 8 3 2 0 0 0 0 |
| − | 8 7 1 0 0 0 0 | + | 8 7 1 0 0 0 0 |
| − | 7 6 2 0 0 0 0 | + | 7 6 2 0 0 0 0 |
| − | 6 5 1 0 0 0 0 | + | 6 5 1 0 0 0 0 |
| − | 5 4 2 0 0 0 0 | + | 5 4 2 0 0 0 0 |
| − | 4 3 1 0 0 0 0 | + | 4 3 1 0 0 0 0 |
| − | 3 9 1 0 0 0 0 | + | 3 9 1 0 0 0 0 |
| − | 9 2 1 0 0 0 0 | + | 9 2 1 0 0 0 0 |
| − | 1 10 2 0 0 0 0 | + | 1 10 2 0 0 0 0 |
| − | 1 55 1 0 0 0 0 | + | 1 55 1 0 0 0 0 |
| − | 2 1 1 0 0 0 0 | + | 2 1 1 0 0 0 0 |
| − | 55 54 1 0 0 0 0 | + | 55 54 1 0 0 0 0 |
| − | S SKP 7 | + | S SKP 7 |
| − | ID BMMCBZ1Sa038 | + | ID BMMCBZ1Sa038 |
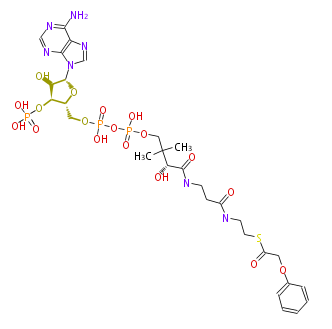
| − | NAME Phenoxy-acetyl-CoA | + | NAME Phenoxy-acetyl-CoA |
| − | FORMULA C29H42N7O18P3S | + | FORMULA C29H42N7O18P3S |
| − | EXACTMASS 901.1519 | + | EXACTMASS 901.1519 |
| − | AVERAGEMASS 901.6674 | + | AVERAGEMASS 901.6674 |
| − | SMILES S(C(COc(c4)cccc4)=O)CCNC(CCNC(=O)[C@@H](C(C)(C)COP(OP(OC[C@H]([C@@H](OP(O)(O)=O)1)O[C@@H](n(c23)cnc2c(ncn3)N)[C@@H]1O)(O)=O)(O)=O)O)=O | + | SMILES S(C(COc(c4)cccc4)=O)CCNC(CCNC(=O)[C@@H](C(C)(C)COP(OP(OC[C@H]([C@@H](OP(O)(O)=O)1)O[C@@H](n(c23)cnc2c(ncn3)N)[C@@H]1O)(O)=O)(O)=O)O)=O |
| − | KEGG http://www.genome.jp/dbget-bin/www_bget?compound+C02577 | + | KEGG http://www.genome.jp/dbget-bin/www_bget?compound+C02577 |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
58 61 0 0 1 0 0 0 0 0999 V2000
6.5817 -3.5672 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.6036 -3.3593 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.9563 -3.8945 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.6473 -4.8456 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.6691 -5.0535 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0000 -4.3103 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.3090 -3.3593 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.2872 -3.1514 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.9344 -4.1024 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
6.8907 -4.5183 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
23.9388 -2.5373 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0
23.0728 -2.0373 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
23.0728 -1.0373 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0
23.9388 -0.5373 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
24.8049 -1.0373 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
24.8049 -2.0373 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
25.5480 -0.3681 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0
25.1413 0.5454 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
24.1467 0.4409 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0
25.6709 -2.5373 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0
23.4776 1.1840 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
23.6855 2.1622 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
22.8195 2.6622 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
22.0764 1.9930 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
21.0982 2.2010 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
24.5991 2.5689 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
22.7150 3.6567 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
22.4831 1.0795 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
20.4291 1.4578 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
23.5240 4.2445 0.0000 P 0 0 0 0 0 0 0 0 0 0 0 0
24.1118 3.4355 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
22.9362 5.0535 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
24.3330 4.8323 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
19.4509 1.6657 0.0000 P 0 0 0 0 0 0 0 0 0 0 0 0
19.2430 0.6876 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
19.6588 2.6439 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
18.4728 1.8736 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
17.8037 1.1305 0.0000 P 0 0 0 0 0 0 0 0 0 0 0 0
18.5468 0.4614 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
17.0605 1.7996 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
17.1345 0.3873 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
16.1564 0.5953 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
15.4872 -0.1479 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
16.2304 -0.8170 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
14.7441 0.5212 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
14.8181 -0.8910 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0
13.8400 -0.6831 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
13.1708 -1.4263 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0
12.1927 -1.2184 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
11.5236 -1.9615 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
10.5454 -1.7536 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
9.8763 -2.4967 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0
8.8981 -2.2888 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
8.2290 -3.0320 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
7.2509 -2.8241 0.0000 S 0 0 0 0 0 0 0 0 0 0 0 0
15.1271 -1.8421 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
13.5309 0.2679 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
10.2364 -0.8025 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
11 16 2 0 0 0 0
11 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
15 16 1 0 0 0 0
15 14 2 0 0 0 0
19 18 1 0 0 0 0
18 17 2 0 0 0 0
17 15 1 0 0 0 0
14 19 1 0 0 0 0
16 20 1 0 0 0 0
24 28 1 6 0 0 0
23 24 1 0 0 0 0
21 28 1 6 0 0 0
23 22 1 0 0 0 0
21 22 1 0 0 0 0
24 25 1 0 0 0 0
25 29 1 0 0 0 0
21 19 1 0 0 0 0
29 34 1 0 0 0 0
34 36 1 0 0 0 0
34 35 2 0 0 0 0
34 37 1 0 0 0 0
41 38 1 0 0 0 0
38 40 2 0 0 0 0
38 39 1 0 0 0 0
41 42 1 0 0 0 0
38 37 1 0 0 0 0
42 43 1 0 0 0 0
43 46 1 0 0 0 0
43 45 1 0 0 0 0
43 44 1 0 0 0 0
46 47 1 0 0 0 0
46 56 1 1 0 0 0
47 57 2 0 0 0 0
47 48 1 0 0 0 0
48 49 1 0 0 0 0
49 50 1 0 0 0 0
50 51 1 0 0 0 0
51 52 1 0 0 0 0
51 58 2 0 0 0 0
52 53 1 0 0 0 0
53 54 1 0 0 0 0
22 26 1 1 0 0 0
23 27 1 1 0 0 0
31 30 1 0 0 0 0
30 33 1 0 0 0 0
30 32 2 0 0 0 0
27 30 1 0 0 0 0
8 3 2 0 0 0 0
8 7 1 0 0 0 0
7 6 2 0 0 0 0
6 5 1 0 0 0 0
5 4 2 0 0 0 0
4 3 1 0 0 0 0
3 9 1 0 0 0 0
9 2 1 0 0 0 0
1 10 2 0 0 0 0
1 55 1 0 0 0 0
2 1 1 0 0 0 0
55 54 1 0 0 0 0
S SKP 7
ID BMMCBZ1Sa038
NAME Phenoxy-acetyl-CoA
FORMULA C29H42N7O18P3S
EXACTMASS 901.1519
AVERAGEMASS 901.6674
SMILES S(C(COc(c4)cccc4)=O)CCNC(CCNC(=O)[C@@H](C(C)(C)COP(OP(OC[C@H]([C@@H](OP(O)(O)=O)1)O[C@@H](n(c23)cnc2c(ncn3)N)[C@@H]1O)(O)=O)(O)=O)O)=O
KEGG http://www.genome.jp/dbget-bin/www_bget?compound+C02577
M END