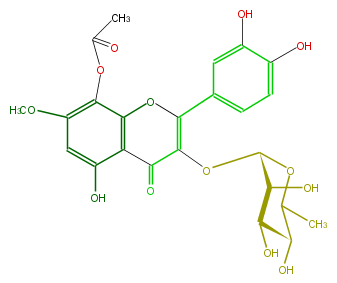
Mol:FL5FFCGS0019
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 37 40 0 0 0 0 0 0 0 0999 V2000 | + | 37 40 0 0 0 0 0 0 0 0999 V2000 |
| − | -2.2503 0.4757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2503 0.4757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.2503 -0.1666 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2503 -0.1666 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6940 -0.4878 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6940 -0.4878 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1377 -0.1666 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1377 -0.1666 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1377 0.4757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1377 0.4757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6940 0.7969 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6940 0.7969 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5814 -0.4878 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5814 -0.4878 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0251 -0.1666 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0251 -0.1666 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0251 0.4757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0251 0.4757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5814 0.7969 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5814 0.7969 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5814 -0.9886 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5814 -0.9886 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6754 0.9205 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6754 0.9205 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2424 0.5932 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2424 0.5932 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8093 0.9205 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8093 0.9205 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8093 1.5752 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8093 1.5752 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2424 1.9026 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2424 1.9026 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6754 1.5752 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6754 1.5752 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6940 -1.1299 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6940 -1.1299 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.4162 1.9256 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.4162 1.9256 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5382 -0.5761 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5382 -0.5761 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2424 2.5571 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2424 2.5571 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.5825 1.4292 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.5825 1.4292 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5909 -1.6287 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 | + | 1.5909 -1.6287 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 |
| − | 2.2191 -1.9914 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.2191 -1.9914 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 2.0198 -1.2939 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 2.0198 -1.2939 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 2.2191 -0.5922 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2191 -0.5922 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5909 -0.2294 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 1.5909 -0.2294 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 1.7902 -0.9269 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 1.7902 -0.9269 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 2.5605 -0.9269 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5605 -0.9269 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.7523 -2.2309 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.7523 -2.2309 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0676 -2.5571 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0676 -2.5571 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.6400 -1.6520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.6400 -1.6520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.7452 2.0367 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.7452 2.0367 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.3098 1.8782 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.3098 1.8782 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.3009 2.4811 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.3009 2.4811 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.9644 0.6286 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.9644 0.6286 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.9422 0.8379 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.9422 0.8379 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 3 18 1 0 0 0 0 | + | 3 18 1 0 0 0 0 |
| − | 19 15 1 0 0 0 0 | + | 19 15 1 0 0 0 0 |
| − | 20 8 1 0 0 0 0 | + | 20 8 1 0 0 0 0 |
| − | 4 3 1 0 0 0 0 | + | 4 3 1 0 0 0 0 |
| − | 16 21 1 0 0 0 0 | + | 16 21 1 0 0 0 0 |
| − | 6 22 1 0 0 0 0 | + | 6 22 1 0 0 0 0 |
| − | 24 23 1 1 0 0 0 | + | 24 23 1 1 0 0 0 |
| − | 24 25 1 0 0 0 0 | + | 24 25 1 0 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 27 1 0 0 0 0 | + | 26 27 1 0 0 0 0 |
| − | 27 28 1 1 0 0 0 | + | 27 28 1 1 0 0 0 |
| − | 28 23 1 1 0 0 0 | + | 28 23 1 1 0 0 0 |
| − | 28 29 1 0 0 0 0 | + | 28 29 1 0 0 0 0 |
| − | 23 30 1 0 0 0 0 | + | 23 30 1 0 0 0 0 |
| − | 24 31 1 0 0 0 0 | + | 24 31 1 0 0 0 0 |
| − | 25 32 1 0 0 0 0 | + | 25 32 1 0 0 0 0 |
| − | 27 20 1 0 0 0 0 | + | 27 20 1 0 0 0 0 |
| − | 22 33 1 0 0 0 0 | + | 22 33 1 0 0 0 0 |
| − | 33 34 2 0 0 0 0 | + | 33 34 2 0 0 0 0 |
| − | 33 35 1 0 0 0 0 | + | 33 35 1 0 0 0 0 |
| − | 1 36 1 0 0 0 0 | + | 1 36 1 0 0 0 0 |
| − | 36 37 1 0 0 0 0 | + | 36 37 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 36 37 | + | M SAL 1 2 36 37 |
| − | M SBL 1 1 39 | + | M SBL 1 1 39 |
| − | M SMT 1 OCH3 | + | M SMT 1 OCH3 |
| − | M SVB 1 39 -2.5922 1.121 | + | M SVB 1 39 -2.5922 1.121 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL5FFCGS0019 | + | ID FL5FFCGS0019 |
| − | KNApSAcK_ID C00006033 | + | KNApSAcK_ID C00006033 |
| − | NAME Purifolin | + | NAME Purifolin |
| − | CAS_RN 62082-22-8 | + | CAS_RN 62082-22-8 |
| − | FORMULA C24H24O13 | + | FORMULA C24H24O13 |
| − | EXACTMASS 520.121690854 | + | EXACTMASS 520.121690854 |
| − | AVERAGEMASS 520.43956 | + | AVERAGEMASS 520.43956 |
| − | SMILES [C@@H]([C@H]1O)(OC(=C(c(c4)cc(c(c4)O)O)2)C(c(c(O)3)c(c(c(OC)c3)OC(C)=O)O2)=O)OC(C)[C@H]([C@H]1O)O | + | SMILES [C@@H]([C@H]1O)(OC(=C(c(c4)cc(c(c4)O)O)2)C(c(c(O)3)c(c(c(OC)c3)OC(C)=O)O2)=O)OC(C)[C@H]([C@H]1O)O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
37 40 0 0 0 0 0 0 0 0999 V2000
-2.2503 0.4757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.2503 -0.1666 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6940 -0.4878 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1377 -0.1666 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1377 0.4757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6940 0.7969 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5814 -0.4878 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.0251 -0.1666 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.0251 0.4757 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5814 0.7969 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.5814 -0.9886 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.6754 0.9205 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2424 0.5932 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8093 0.9205 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8093 1.5752 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2424 1.9026 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.6754 1.5752 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6940 -1.1299 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.4162 1.9256 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.5382 -0.5761 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.2424 2.5571 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.5825 1.4292 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.5909 -1.6287 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0
2.2191 -1.9914 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
2.0198 -1.2939 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
2.2191 -0.5922 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.5909 -0.2294 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
1.7902 -0.9269 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
2.5605 -0.9269 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.7523 -2.2309 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.0676 -2.5571 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.6400 -1.6520 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.7452 2.0367 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.3098 1.8782 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.3009 2.4811 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.9644 0.6286 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.9422 0.8379 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
3 18 1 0 0 0 0
19 15 1 0 0 0 0
20 8 1 0 0 0 0
4 3 1 0 0 0 0
16 21 1 0 0 0 0
6 22 1 0 0 0 0
24 23 1 1 0 0 0
24 25 1 0 0 0 0
25 26 1 0 0 0 0
26 27 1 0 0 0 0
27 28 1 1 0 0 0
28 23 1 1 0 0 0
28 29 1 0 0 0 0
23 30 1 0 0 0 0
24 31 1 0 0 0 0
25 32 1 0 0 0 0
27 20 1 0 0 0 0
22 33 1 0 0 0 0
33 34 2 0 0 0 0
33 35 1 0 0 0 0
1 36 1 0 0 0 0
36 37 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 36 37
M SBL 1 1 39
M SMT 1 OCH3
M SVB 1 39 -2.5922 1.121
S SKP 8
ID FL5FFCGS0019
KNApSAcK_ID C00006033
NAME Purifolin
CAS_RN 62082-22-8
FORMULA C24H24O13
EXACTMASS 520.121690854
AVERAGEMASS 520.43956
SMILES [C@@H]([C@H]1O)(OC(=C(c(c4)cc(c(c4)O)O)2)C(c(c(O)3)c(c(c(OC)c3)OC(C)=O)O2)=O)OC(C)[C@H]([C@H]1O)O
M END