Mol:FL5FACGA0031
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 44 48 0 0 0 0 0 0 0 0999 V2000 | + | 44 48 0 0 0 0 0 0 0 0999 V2000 |
| − | -2.7968 0.8678 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7968 0.8678 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7968 0.2254 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7968 0.2254 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.2405 -0.0958 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2405 -0.0958 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6842 0.2254 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6842 0.2254 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6842 0.8678 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6842 0.8678 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.2405 1.1889 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2405 1.1889 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1279 -0.0958 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1279 -0.0958 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5716 0.2254 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5716 0.2254 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5716 0.8678 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5716 0.8678 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1279 1.1889 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1279 1.1889 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1279 -0.5966 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1279 -0.5966 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0155 1.1888 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0155 1.1888 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5514 0.8615 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5514 0.8615 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1184 1.1888 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1184 1.1888 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1184 1.8435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1184 1.8435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5514 2.1708 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5514 2.1708 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0155 1.8435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0155 1.8435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.2405 -0.7379 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2405 -0.7379 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.9121 2.2228 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.9121 2.2228 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.2918 1.1889 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.2918 1.1889 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0422 -0.1290 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0422 -0.1290 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5514 2.8253 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5514 2.8253 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8143 0.3509 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8143 0.3509 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2567 -0.2331 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2567 -0.2331 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8938 0.0147 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8938 0.0147 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.6729 0.0634 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.6729 0.0634 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0618 0.4681 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0618 0.4681 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5045 0.1739 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5045 0.1739 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5670 -0.2666 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5670 -0.2666 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2588 -0.5981 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2588 -0.5981 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.6729 -0.6406 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.6729 -0.6406 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5670 -1.0955 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5670 -1.0955 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.0671 -1.3842 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.0671 -1.3842 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1946 -1.3105 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1946 -1.3105 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.0671 -1.9609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.0671 -1.9609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5665 -2.2492 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5665 -2.2492 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0659 -1.9609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0659 -1.9609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0659 -1.3842 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0659 -1.3842 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5665 -1.0959 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5665 -1.0959 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5665 -2.8253 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5665 -2.8253 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.5648 -2.2490 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5648 -2.2490 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.5648 -1.0962 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5648 -1.0962 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.5773 0.4957 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5773 0.4957 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.2918 0.0832 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.2918 0.0832 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 3 18 1 0 0 0 0 | + | 3 18 1 0 0 0 0 |
| − | 19 15 1 0 0 0 0 | + | 19 15 1 0 0 0 0 |
| − | 20 1 1 0 0 0 0 | + | 20 1 1 0 0 0 0 |
| − | 21 8 1 0 0 0 0 | + | 21 8 1 0 0 0 0 |
| − | 16 22 1 0 0 0 0 | + | 16 22 1 0 0 0 0 |
| − | 23 24 1 1 0 0 0 | + | 23 24 1 1 0 0 0 |
| − | 24 25 1 1 0 0 0 | + | 24 25 1 1 0 0 0 |
| − | 26 25 1 1 0 0 0 | + | 26 25 1 1 0 0 0 |
| − | 26 27 1 0 0 0 0 | + | 26 27 1 0 0 0 0 |
| − | 27 28 1 0 0 0 0 | + | 27 28 1 0 0 0 0 |
| − | 28 23 1 0 0 0 0 | + | 28 23 1 0 0 0 0 |
| − | 24 29 1 0 0 0 0 | + | 24 29 1 0 0 0 0 |
| − | 25 30 1 0 0 0 0 | + | 25 30 1 0 0 0 0 |
| − | 21 23 1 0 0 0 0 | + | 21 23 1 0 0 0 0 |
| − | 26 31 1 0 0 0 0 | + | 26 31 1 0 0 0 0 |
| − | 29 32 1 0 0 0 0 | + | 29 32 1 0 0 0 0 |
| − | 32 33 1 0 0 0 0 | + | 32 33 1 0 0 0 0 |
| − | 32 34 2 0 0 0 0 | + | 32 34 2 0 0 0 0 |
| − | 33 35 2 0 0 0 0 | + | 33 35 2 0 0 0 0 |
| − | 35 36 1 0 0 0 0 | + | 35 36 1 0 0 0 0 |
| − | 36 37 2 0 0 0 0 | + | 36 37 2 0 0 0 0 |
| − | 37 38 1 0 0 0 0 | + | 37 38 1 0 0 0 0 |
| − | 38 39 2 0 0 0 0 | + | 38 39 2 0 0 0 0 |
| − | 39 33 1 0 0 0 0 | + | 39 33 1 0 0 0 0 |
| − | 36 40 1 0 0 0 0 | + | 36 40 1 0 0 0 0 |
| − | 37 41 1 0 0 0 0 | + | 37 41 1 0 0 0 0 |
| − | 38 42 1 0 0 0 0 | + | 38 42 1 0 0 0 0 |
| − | 27 43 1 0 0 0 0 | + | 27 43 1 0 0 0 0 |
| − | 43 44 1 0 0 0 0 | + | 43 44 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 43 44 | + | M SAL 1 2 43 44 |
| − | M SBL 1 1 47 | + | M SBL 1 1 47 |
| − | M SMT 1 CH2OH | + | M SMT 1 CH2OH |
| − | M SBV 1 47 -4.7109 3.5766 | + | M SBV 1 47 -4.7109 3.5766 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL5FACGA0031 | + | ID FL5FACGA0031 |
| − | KNApSAcK_ID C00005950 | + | KNApSAcK_ID C00005950 |
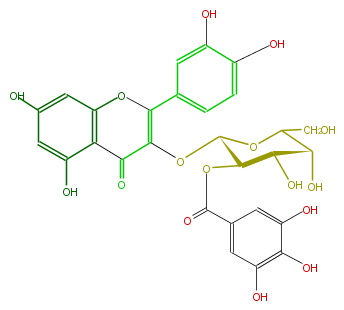
| − | NAME Quercetin 3-(2''-galloylgalactoside) | + | NAME Quercetin 3-(2''-galloylgalactoside) |
| − | CAS_RN - | + | CAS_RN - |
| − | FORMULA C28H24O16 | + | FORMULA C28H24O16 |
| − | EXACTMASS 616.1064347199999 | + | EXACTMASS 616.1064347199999 |
| − | AVERAGEMASS 616.48056 | + | AVERAGEMASS 616.48056 |
| − | SMILES O=C(c(c5)cc(O)c(O)c5O)OC(C4O)C(OC(CO)C4O)OC(C(=O)1)=C(c(c3)cc(c(c3)O)O)Oc(c2)c(c(cc(O)2)O)1 | + | SMILES O=C(c(c5)cc(O)c(O)c5O)OC(C4O)C(OC(CO)C4O)OC(C(=O)1)=C(c(c3)cc(c(c3)O)O)Oc(c2)c(c(cc(O)2)O)1 |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
44 48 0 0 0 0 0 0 0 0999 V2000
-2.7968 0.8678 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7968 0.2254 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.2405 -0.0958 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6842 0.2254 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6842 0.8678 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.2405 1.1889 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1279 -0.0958 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5716 0.2254 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5716 0.8678 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1279 1.1889 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.1279 -0.5966 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.0155 1.1888 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5514 0.8615 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1184 1.1888 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1184 1.8435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5514 2.1708 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.0155 1.8435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.2405 -0.7379 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.9121 2.2228 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.2918 1.1889 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.0422 -0.1290 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.5514 2.8253 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.8143 0.3509 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2567 -0.2331 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8938 0.0147 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.6729 0.0634 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0618 0.4681 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.5045 0.1739 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.5670 -0.2666 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.2588 -0.5981 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.6729 -0.6406 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.5670 -1.0955 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.0671 -1.3842 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.1946 -1.3105 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.0671 -1.9609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.5665 -2.2492 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0659 -1.9609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0659 -1.3842 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.5665 -1.0959 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.5665 -2.8253 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.5648 -2.2490 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.5648 -1.0962 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.5773 0.4957 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.2918 0.0832 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
3 18 1 0 0 0 0
19 15 1 0 0 0 0
20 1 1 0 0 0 0
21 8 1 0 0 0 0
16 22 1 0 0 0 0
23 24 1 1 0 0 0
24 25 1 1 0 0 0
26 25 1 1 0 0 0
26 27 1 0 0 0 0
27 28 1 0 0 0 0
28 23 1 0 0 0 0
24 29 1 0 0 0 0
25 30 1 0 0 0 0
21 23 1 0 0 0 0
26 31 1 0 0 0 0
29 32 1 0 0 0 0
32 33 1 0 0 0 0
32 34 2 0 0 0 0
33 35 2 0 0 0 0
35 36 1 0 0 0 0
36 37 2 0 0 0 0
37 38 1 0 0 0 0
38 39 2 0 0 0 0
39 33 1 0 0 0 0
36 40 1 0 0 0 0
37 41 1 0 0 0 0
38 42 1 0 0 0 0
27 43 1 0 0 0 0
43 44 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 43 44
M SBL 1 1 47
M SMT 1 CH2OH
M SBV 1 47 -4.7109 3.5766
S SKP 8
ID FL5FACGA0031
KNApSAcK_ID C00005950
NAME Quercetin 3-(2''-galloylgalactoside)
CAS_RN -
FORMULA C28H24O16
EXACTMASS 616.1064347199999
AVERAGEMASS 616.48056
SMILES O=C(c(c5)cc(O)c(O)c5O)OC(C4O)C(OC(CO)C4O)OC(C(=O)1)=C(c(c3)cc(c(c3)O)O)Oc(c2)c(c(cc(O)2)O)1
M END