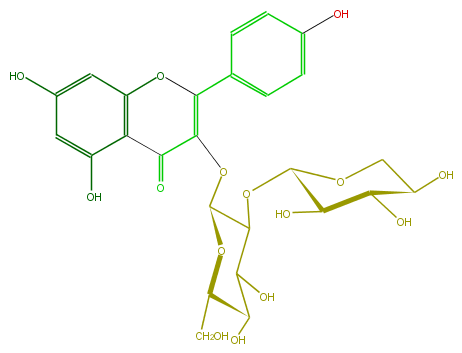
Mol:FL5FAAGL0005
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 41 45 0 0 0 0 0 0 0 0999 V2000 | + | 41 45 0 0 0 0 0 0 0 0999 V2000 |
| − | -3.5194 1.6227 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.5194 1.6227 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.5194 0.8133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.5194 0.8133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.8184 0.4085 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.8184 0.4085 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1174 0.8133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1174 0.8133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1174 1.6227 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1174 1.6227 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.8184 2.0274 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.8184 2.0274 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4164 0.4085 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4164 0.4085 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.7154 0.8133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7154 0.8133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.7154 1.6227 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7154 1.6227 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4164 2.0274 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4164 2.0274 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4164 -0.2226 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4164 -0.2226 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0146 2.0273 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0146 2.0273 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6999 1.6148 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6999 1.6148 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.4144 2.0273 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.4144 2.0273 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.4144 2.8523 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.4144 2.8523 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6999 3.2648 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6999 3.2648 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0146 2.8523 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0146 2.8523 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.8184 -0.4006 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.8184 -0.4006 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.2202 2.0273 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.2202 2.0273 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.1287 3.2646 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.1287 3.2646 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.4492 -2.3609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.4492 -2.3609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.3426 -2.8180 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.3426 -2.8180 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0913 -1.9390 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0913 -1.9390 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.3426 -1.0548 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.3426 -1.0548 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.4492 -0.5976 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.4492 -0.5976 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.1980 -1.4765 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.1980 -1.4765 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.1604 0.0951 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.1604 0.0951 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.3050 -0.3363 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.3050 -0.3363 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6336 -2.4002 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6336 -2.4002 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1685 0.1619 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1685 0.1619 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8182 -0.6958 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8182 -0.6958 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.7539 -0.3320 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.7539 -0.3320 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.6566 -0.3222 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.6566 -0.3222 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0006 0.3340 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0006 0.3340 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.1820 -0.0980 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.1820 -0.0980 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.0954 -0.7258 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.0954 -0.7258 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.3738 -0.9034 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.3738 -0.9034 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.2202 0.0430 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.2202 0.0430 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0684 -3.2648 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0684 -3.2648 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6429 -3.1803 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6429 -3.1803 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1627 -2.2801 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1627 -2.2801 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 3 18 1 0 0 0 0 | + | 3 18 1 0 0 0 0 |
| − | 1 19 1 0 0 0 0 | + | 1 19 1 0 0 0 0 |
| − | 15 20 1 0 0 0 0 | + | 15 20 1 0 0 0 0 |
| − | 22 21 1 1 0 0 0 | + | 22 21 1 1 0 0 0 |
| − | 22 23 1 0 0 0 0 | + | 22 23 1 0 0 0 0 |
| − | 23 24 1 0 0 0 0 | + | 23 24 1 0 0 0 0 |
| − | 24 25 1 0 0 0 0 | + | 24 25 1 0 0 0 0 |
| − | 25 26 1 1 0 0 0 | + | 25 26 1 1 0 0 0 |
| − | 26 21 1 1 0 0 0 | + | 26 21 1 1 0 0 0 |
| − | 25 27 1 0 0 0 0 | + | 25 27 1 0 0 0 0 |
| − | 24 28 1 0 0 0 0 | + | 24 28 1 0 0 0 0 |
| − | 23 29 1 0 0 0 0 | + | 23 29 1 0 0 0 0 |
| − | 30 31 1 1 0 0 0 | + | 30 31 1 1 0 0 0 |
| − | 31 32 1 1 0 0 0 | + | 31 32 1 1 0 0 0 |
| − | 33 32 1 1 0 0 0 | + | 33 32 1 1 0 0 0 |
| − | 33 34 1 0 0 0 0 | + | 33 34 1 0 0 0 0 |
| − | 34 35 1 0 0 0 0 | + | 34 35 1 0 0 0 0 |
| − | 35 30 1 0 0 0 0 | + | 35 30 1 0 0 0 0 |
| − | 31 36 1 0 0 0 0 | + | 31 36 1 0 0 0 0 |
| − | 32 37 1 0 0 0 0 | + | 32 37 1 0 0 0 0 |
| − | 30 28 1 0 0 0 0 | + | 30 28 1 0 0 0 0 |
| − | 33 38 1 0 0 0 0 | + | 33 38 1 0 0 0 0 |
| − | 22 39 1 0 0 0 0 | + | 22 39 1 0 0 0 0 |
| − | 27 8 1 0 0 0 0 | + | 27 8 1 0 0 0 0 |
| − | 40 41 1 0 0 0 0 | + | 40 41 1 0 0 0 0 |
| − | 21 40 1 0 0 0 0 | + | 21 40 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 40 41 | + | M SAL 1 2 40 41 |
| − | M SBL 1 1 45 | + | M SBL 1 1 45 |
| − | M SMT 1 ^CH2OH | + | M SMT 1 ^CH2OH |
| − | M SBV 1 45 0.1937 0.8194 | + | M SBV 1 45 0.1937 0.8194 |
| − | S SKP 5 | + | S SKP 5 |
| − | ID FL5FAAGL0005 | + | ID FL5FAAGL0005 |
| − | FORMULA C26H28O15 | + | FORMULA C26H28O15 |
| − | EXACTMASS 580.1428202259999 | + | EXACTMASS 580.1428202259999 |
| − | AVERAGEMASS 580.49152 | + | AVERAGEMASS 580.49152 |
| − | SMILES c(O4)(c(C(C(=C4c(c5)ccc(c5)O)OC(O2)C(OC(C3O)OCC(C3O)O)C(O)C(C2CO)O)=O)1)cc(O)cc1O | + | SMILES c(O4)(c(C(C(=C4c(c5)ccc(c5)O)OC(O2)C(OC(C3O)OCC(C3O)O)C(O)C(C2CO)O)=O)1)cc(O)cc1O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
41 45 0 0 0 0 0 0 0 0999 V2000
-3.5194 1.6227 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.5194 0.8133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.8184 0.4085 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.1174 0.8133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.1174 1.6227 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.8184 2.0274 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.4164 0.4085 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.7154 0.8133 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.7154 1.6227 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.4164 2.0274 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.4164 -0.2226 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.0146 2.0273 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.6999 1.6148 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.4144 2.0273 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.4144 2.8523 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.6999 3.2648 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.0146 2.8523 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.8184 -0.4006 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.2202 2.0273 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.1287 3.2646 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.4492 -2.3609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.3426 -2.8180 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0913 -1.9390 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.3426 -1.0548 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.4492 -0.5976 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.1980 -1.4765 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.1604 0.0951 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.3050 -0.3363 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.6336 -2.4002 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.1685 0.1619 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8182 -0.6958 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.7539 -0.3320 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.6566 -0.3222 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.0006 0.3340 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.1820 -0.0980 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.0954 -0.7258 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.3738 -0.9034 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.2202 0.0430 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.0684 -3.2648 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.6429 -3.1803 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1627 -2.2801 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
3 18 1 0 0 0 0
1 19 1 0 0 0 0
15 20 1 0 0 0 0
22 21 1 1 0 0 0
22 23 1 0 0 0 0
23 24 1 0 0 0 0
24 25 1 0 0 0 0
25 26 1 1 0 0 0
26 21 1 1 0 0 0
25 27 1 0 0 0 0
24 28 1 0 0 0 0
23 29 1 0 0 0 0
30 31 1 1 0 0 0
31 32 1 1 0 0 0
33 32 1 1 0 0 0
33 34 1 0 0 0 0
34 35 1 0 0 0 0
35 30 1 0 0 0 0
31 36 1 0 0 0 0
32 37 1 0 0 0 0
30 28 1 0 0 0 0
33 38 1 0 0 0 0
22 39 1 0 0 0 0
27 8 1 0 0 0 0
40 41 1 0 0 0 0
21 40 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 40 41
M SBL 1 1 45
M SMT 1 ^CH2OH
M SBV 1 45 0.1937 0.8194
S SKP 5
ID FL5FAAGL0005
FORMULA C26H28O15
EXACTMASS 580.1428202259999
AVERAGEMASS 580.49152
SMILES c(O4)(c(C(C(=C4c(c5)ccc(c5)O)OC(O2)C(OC(C3O)OCC(C3O)O)C(O)C(C2CO)O)=O)1)cc(O)cc1O
M END