Mol:FL3FACDS0017
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 55 60 0 0 0 0 0 0 0 0999 V2000 | + | 55 60 0 0 0 0 0 0 0 0999 V2000 |
| − | -1.6593 0.2058 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6593 0.2058 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6593 -0.4366 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6593 -0.4366 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1030 -0.7578 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1030 -0.7578 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5467 -0.4366 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5467 -0.4366 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5467 0.2058 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5467 0.2058 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1030 0.5269 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1030 0.5269 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0096 -0.7578 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0096 -0.7578 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5659 -0.4366 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5659 -0.4366 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5659 0.2058 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5659 0.2058 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0096 0.5269 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0096 0.5269 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0096 -1.2586 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0096 -1.2586 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.2154 0.5268 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2154 0.5268 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1030 -1.3999 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1030 -1.3999 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2458 0.6299 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2458 0.6299 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8320 0.2915 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8320 0.2915 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.4182 0.6299 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.4182 0.6299 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.4182 1.3069 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.4182 1.3069 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8320 1.6453 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8320 1.6453 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2458 1.3069 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2458 1.3069 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0041 1.6451 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0041 1.6451 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.2468 4.2427 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.2468 4.2427 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.6412 3.7839 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.6412 3.7839 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.8981 3.1232 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.8981 3.1232 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.9050 2.4858 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.9050 2.4858 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.3683 2.9490 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.3683 2.9490 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0633 3.5270 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0633 3.5270 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.7788 5.0532 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.7788 5.0532 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.6064 4.4991 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.6064 4.4991 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2627 2.7447 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2627 2.7447 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.8392 -0.6635 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.8392 -0.6635 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.4680 -1.1534 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.4680 -1.1534 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.9336 -0.9456 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.9336 -0.9456 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.4178 -0.9400 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.4178 -0.9400 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7926 -0.5651 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7926 -0.5651 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.2602 -0.8119 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.2602 -0.8119 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.3531 -0.9601 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.3531 -0.9601 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.0467 -1.1816 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.0467 -1.1816 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.6273 -1.4597 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.6273 -1.4597 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.6273 -2.1279 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.6273 -2.1279 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.2766 -2.3018 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.2766 -2.3018 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1439 -2.4070 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1439 -2.4070 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1439 -3.0943 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1439 -3.0943 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4269 -3.4449 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4269 -3.4449 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4269 -4.0886 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4269 -4.0886 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.8693 -4.4105 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.8693 -4.4105 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.3118 -4.0886 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.3118 -4.0886 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.3118 -3.4449 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.3118 -3.4449 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.8693 -3.1230 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.8693 -3.1230 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.2448 -4.4100 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.2448 -4.4100 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.8693 -5.0532 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.8693 -5.0532 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.8686 0.3699 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.8686 0.3699 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.6386 3.8591 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.6386 3.8591 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.3531 3.4466 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.3531 3.4466 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.6586 -0.0278 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.6586 -0.0278 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.8617 -0.2413 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.8617 -0.2413 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 1 12 1 0 0 0 0 | + | 1 12 1 0 0 0 0 |
| − | 3 13 1 0 0 0 0 | + | 3 13 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 18 1 0 0 0 0 | + | 17 18 1 0 0 0 0 |
| − | 18 19 2 0 0 0 0 | + | 18 19 2 0 0 0 0 |
| − | 19 14 1 0 0 0 0 | + | 19 14 1 0 0 0 0 |
| − | 9 14 1 0 0 0 0 | + | 9 14 1 0 0 0 0 |
| − | 17 20 1 0 0 0 0 | + | 17 20 1 0 0 0 0 |
| − | 21 22 1 1 0 0 0 | + | 21 22 1 1 0 0 0 |
| − | 22 23 1 1 0 0 0 | + | 22 23 1 1 0 0 0 |
| − | 24 23 1 1 0 0 0 | + | 24 23 1 1 0 0 0 |
| − | 24 25 1 0 0 0 0 | + | 24 25 1 0 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 21 1 0 0 0 0 | + | 26 21 1 0 0 0 0 |
| − | 21 27 1 0 0 0 0 | + | 21 27 1 0 0 0 0 |
| − | 22 28 1 0 0 0 0 | + | 22 28 1 0 0 0 0 |
| − | 23 29 1 0 0 0 0 | + | 23 29 1 0 0 0 0 |
| − | 20 24 1 0 0 0 0 | + | 20 24 1 0 0 0 0 |
| − | 30 31 1 1 0 0 0 | + | 30 31 1 1 0 0 0 |
| − | 31 32 1 1 0 0 0 | + | 31 32 1 1 0 0 0 |
| − | 33 32 1 1 0 0 0 | + | 33 32 1 1 0 0 0 |
| − | 33 34 1 0 0 0 0 | + | 33 34 1 0 0 0 0 |
| − | 34 35 1 0 0 0 0 | + | 34 35 1 0 0 0 0 |
| − | 35 30 1 0 0 0 0 | + | 35 30 1 0 0 0 0 |
| − | 30 36 1 0 0 0 0 | + | 30 36 1 0 0 0 0 |
| − | 31 37 1 0 0 0 0 | + | 31 37 1 0 0 0 0 |
| − | 32 38 1 0 0 0 0 | + | 32 38 1 0 0 0 0 |
| − | 2 33 1 0 0 0 0 | + | 2 33 1 0 0 0 0 |
| − | 38 39 1 0 0 0 0 | + | 38 39 1 0 0 0 0 |
| − | 39 40 2 0 0 0 0 | + | 39 40 2 0 0 0 0 |
| − | 39 41 1 0 0 0 0 | + | 39 41 1 0 0 0 0 |
| − | 41 42 2 0 0 0 0 | + | 41 42 2 0 0 0 0 |
| − | 42 43 1 0 0 0 0 | + | 42 43 1 0 0 0 0 |
| − | 43 44 2 0 0 0 0 | + | 43 44 2 0 0 0 0 |
| − | 44 45 1 0 0 0 0 | + | 44 45 1 0 0 0 0 |
| − | 45 46 2 0 0 0 0 | + | 45 46 2 0 0 0 0 |
| − | 46 47 1 0 0 0 0 | + | 46 47 1 0 0 0 0 |
| − | 47 48 2 0 0 0 0 | + | 47 48 2 0 0 0 0 |
| − | 48 43 1 0 0 0 0 | + | 48 43 1 0 0 0 0 |
| − | 46 49 1 0 0 0 0 | + | 46 49 1 0 0 0 0 |
| − | 45 50 1 0 0 0 0 | + | 45 50 1 0 0 0 0 |
| − | 16 51 1 0 0 0 0 | + | 16 51 1 0 0 0 0 |
| − | 26 52 1 0 0 0 0 | + | 26 52 1 0 0 0 0 |
| − | 52 53 1 0 0 0 0 | + | 52 53 1 0 0 0 0 |
| − | 35 54 1 0 0 0 0 | + | 35 54 1 0 0 0 0 |
| − | 54 55 1 0 0 0 0 | + | 54 55 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 52 53 | + | M SAL 1 2 52 53 |
| − | M SBL 1 1 57 | + | M SBL 1 1 57 |
| − | M SMT 1 CH2OH | + | M SMT 1 CH2OH |
| − | M SBV 1 57 -6.1330 7.1379 | + | M SBV 1 57 -6.1330 7.1379 |
| − | M STY 1 2 SUP | + | M STY 1 2 SUP |
| − | M SLB 1 2 2 | + | M SLB 1 2 2 |
| − | M SAL 2 2 54 55 | + | M SAL 2 2 54 55 |
| − | M SBL 2 1 59 | + | M SBL 2 1 59 |
| − | M SMT 2 ^CH2OH | + | M SMT 2 ^CH2OH |
| − | M SBV 2 59 -7.1067 7.5900 | + | M SBV 2 59 -7.1067 7.5900 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL3FACDS0017 | + | ID FL3FACDS0017 |
| − | KNApSAcK_ID C00006330 | + | KNApSAcK_ID C00006330 |
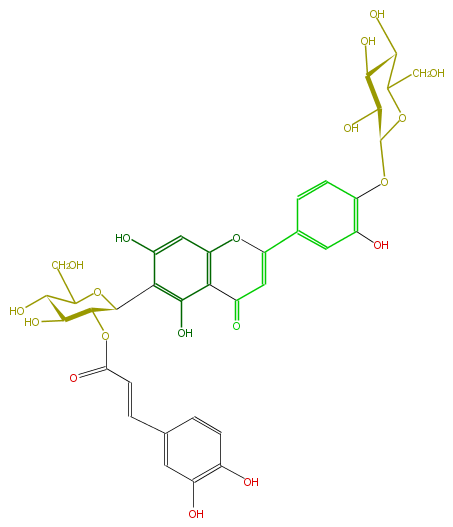
| − | NAME Isoorientin 4'-O-glucoside-2''-O-(E)-caffeate | + | NAME Isoorientin 4'-O-glucoside-2''-O-(E)-caffeate |
| − | CAS_RN 64763-01-5 | + | CAS_RN 64763-01-5 |
| − | FORMULA C36H36O19 | + | FORMULA C36H36O19 |
| − | EXACTMASS 772.18507897 | + | EXACTMASS 772.18507897 |
| − | AVERAGEMASS 772.6596400000001 | + | AVERAGEMASS 772.6596400000001 |
| − | SMILES OC(C(CO)6)C(O)C(C(O6)Oc(c1)c(cc(C(=C2)Oc(c5)c(c(O)c(c5O)C(C(OC(C=Cc(c4)ccc(O)c(O)4)=O)3)OC(C(C(O)3)O)CO)C2=O)c1)O)O | + | SMILES OC(C(CO)6)C(O)C(C(O6)Oc(c1)c(cc(C(=C2)Oc(c5)c(c(O)c(c5O)C(C(OC(C=Cc(c4)ccc(O)c(O)4)=O)3)OC(C(C(O)3)O)CO)C2=O)c1)O)O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
55 60 0 0 0 0 0 0 0 0999 V2000
-1.6593 0.2058 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6593 -0.4366 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1030 -0.7578 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5467 -0.4366 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5467 0.2058 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1030 0.5269 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0096 -0.7578 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5659 -0.4366 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5659 0.2058 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0096 0.5269 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.0096 -1.2586 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.2154 0.5268 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.1030 -1.3999 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.2458 0.6299 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8320 0.2915 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.4182 0.6299 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.4182 1.3069 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8320 1.6453 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.2458 1.3069 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.0041 1.6451 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.2468 4.2427 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.6412 3.7839 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.8981 3.1232 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.9050 2.4858 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.3683 2.9490 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.0633 3.5270 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.7788 5.0532 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.6064 4.4991 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.2627 2.7447 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.8392 -0.6635 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.4680 -1.1534 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.9336 -0.9456 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.4178 -0.9400 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7926 -0.5651 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.2602 -0.8119 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.3531 -0.9601 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.0467 -1.1816 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.6273 -1.4597 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.6273 -2.1279 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.2766 -2.3018 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.1439 -2.4070 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.1439 -3.0943 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.4269 -3.4449 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.4269 -4.0886 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.8693 -4.4105 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.3118 -4.0886 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.3118 -3.4449 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.8693 -3.1230 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.2448 -4.4100 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.8693 -5.0532 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.8686 0.3699 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.6386 3.8591 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3531 3.4466 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.6586 -0.0278 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.8617 -0.2413 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
1 12 1 0 0 0 0
3 13 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 18 1 0 0 0 0
18 19 2 0 0 0 0
19 14 1 0 0 0 0
9 14 1 0 0 0 0
17 20 1 0 0 0 0
21 22 1 1 0 0 0
22 23 1 1 0 0 0
24 23 1 1 0 0 0
24 25 1 0 0 0 0
25 26 1 0 0 0 0
26 21 1 0 0 0 0
21 27 1 0 0 0 0
22 28 1 0 0 0 0
23 29 1 0 0 0 0
20 24 1 0 0 0 0
30 31 1 1 0 0 0
31 32 1 1 0 0 0
33 32 1 1 0 0 0
33 34 1 0 0 0 0
34 35 1 0 0 0 0
35 30 1 0 0 0 0
30 36 1 0 0 0 0
31 37 1 0 0 0 0
32 38 1 0 0 0 0
2 33 1 0 0 0 0
38 39 1 0 0 0 0
39 40 2 0 0 0 0
39 41 1 0 0 0 0
41 42 2 0 0 0 0
42 43 1 0 0 0 0
43 44 2 0 0 0 0
44 45 1 0 0 0 0
45 46 2 0 0 0 0
46 47 1 0 0 0 0
47 48 2 0 0 0 0
48 43 1 0 0 0 0
46 49 1 0 0 0 0
45 50 1 0 0 0 0
16 51 1 0 0 0 0
26 52 1 0 0 0 0
52 53 1 0 0 0 0
35 54 1 0 0 0 0
54 55 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 52 53
M SBL 1 1 57
M SMT 1 CH2OH
M SBV 1 57 -6.1330 7.1379
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 54 55
M SBL 2 1 59
M SMT 2 ^CH2OH
M SBV 2 59 -7.1067 7.5900
S SKP 8
ID FL3FACDS0017
KNApSAcK_ID C00006330
NAME Isoorientin 4'-O-glucoside-2''-O-(E)-caffeate
CAS_RN 64763-01-5
FORMULA C36H36O19
EXACTMASS 772.18507897
AVERAGEMASS 772.6596400000001
SMILES OC(C(CO)6)C(O)C(C(O6)Oc(c1)c(cc(C(=C2)Oc(c5)c(c(O)c(c5O)C(C(OC(C=Cc(c4)ccc(O)c(O)4)=O)3)OC(C(C(O)3)O)CO)C2=O)c1)O)O
M END