Mol:FL3FACCS0046
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 45 49 0 0 0 0 0 0 0 0999 V2000 | + | 45 49 0 0 0 0 0 0 0 0999 V2000 |
| − | -0.6182 1.7612 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6182 1.7612 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0484 2.1460 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0484 2.1460 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0484 2.9157 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0484 2.9157 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6182 3.3005 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6182 3.3005 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.2847 2.9157 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.2847 2.9157 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.2847 2.1460 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.2847 2.1460 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7150 1.7612 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7150 1.7612 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3815 2.1460 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3815 2.1460 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3815 2.9157 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3815 2.9157 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7150 3.3005 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7150 3.3005 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0199 1.7774 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0199 1.7774 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0199 1.0468 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0199 1.0468 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.6527 0.6814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.6527 0.6814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.2855 1.0468 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.2855 1.0468 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.2855 1.7774 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.2855 1.7774 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.6527 2.1428 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.6527 2.1428 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.7200 0.7959 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.7200 0.7959 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7150 3.7581 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7150 3.7581 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6182 3.7603 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6182 3.7603 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.8125 1.8413 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.8125 1.8413 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.2941 1.0682 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.2941 1.0682 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7543 0.4443 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7543 0.4443 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.0100 0.8001 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.0100 0.8001 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1882 0.7269 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1882 0.7269 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.7279 1.3509 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.7279 1.3509 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.4724 0.9951 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.4724 0.9951 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.7960 0.5663 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.7960 0.5663 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.3688 0.6089 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.3688 0.6089 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6943 0.2533 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6943 0.2533 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7953 1.5544 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7953 1.5544 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.1956 1.7855 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.1956 1.7855 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1920 -0.2489 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1920 -0.2489 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6245 0.0787 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6245 0.0787 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1920 -0.8293 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1920 -0.8293 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.7960 2.0722 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.7960 2.0722 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.8156 -1.8621 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.8156 -1.8621 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.4012 -2.2002 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.4012 -2.2002 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.4012 -2.8764 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.4012 -2.8764 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.8156 -3.2145 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.8156 -3.2145 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.2299 -2.8764 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.2299 -2.8764 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.2299 -2.2002 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.2299 -2.2002 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.8156 -3.7603 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.8156 -3.7603 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.8942 -3.1610 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.8942 -3.1610 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.6345 -3.1610 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.6345 -3.1610 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.8019 -1.1815 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.8019 -1.1815 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 2 0 0 0 0 | + | 1 2 2 0 0 0 0 |
| − | 2 3 1 0 0 0 0 | + | 2 3 1 0 0 0 0 |
| − | 3 4 2 0 0 0 0 | + | 3 4 2 0 0 0 0 |
| − | 4 5 1 0 0 0 0 | + | 4 5 1 0 0 0 0 |
| − | 5 6 2 0 0 0 0 | + | 5 6 2 0 0 0 0 |
| − | 6 1 1 0 0 0 0 | + | 6 1 1 0 0 0 0 |
| − | 2 7 1 0 0 0 0 | + | 2 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 3 1 0 0 0 0 | + | 10 3 1 0 0 0 0 |
| − | 8 11 1 0 0 0 0 | + | 8 11 1 0 0 0 0 |
| − | 11 12 2 0 0 0 0 | + | 11 12 2 0 0 0 0 |
| − | 12 13 1 0 0 0 0 | + | 12 13 1 0 0 0 0 |
| − | 13 14 2 0 0 0 0 | + | 13 14 2 0 0 0 0 |
| − | 14 15 1 0 0 0 0 | + | 14 15 1 0 0 0 0 |
| − | 15 16 2 0 0 0 0 | + | 15 16 2 0 0 0 0 |
| − | 16 11 1 0 0 0 0 | + | 16 11 1 0 0 0 0 |
| − | 14 17 1 0 0 0 0 | + | 14 17 1 0 0 0 0 |
| − | 10 18 2 0 0 0 0 | + | 10 18 2 0 0 0 0 |
| − | 4 19 1 0 0 0 0 | + | 4 19 1 0 0 0 0 |
| − | 6 20 1 0 0 0 0 | + | 6 20 1 0 0 0 0 |
| − | 21 22 1 1 0 0 0 | + | 21 22 1 1 0 0 0 |
| − | 22 23 1 1 0 0 0 | + | 22 23 1 1 0 0 0 |
| − | 24 23 1 1 0 0 0 | + | 24 23 1 1 0 0 0 |
| − | 24 25 1 0 0 0 0 | + | 24 25 1 0 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 21 1 0 0 0 0 | + | 26 21 1 0 0 0 0 |
| − | 1 24 1 0 0 0 0 | + | 1 24 1 0 0 0 0 |
| − | 21 27 1 0 0 0 0 | + | 21 27 1 0 0 0 0 |
| − | 22 28 1 0 0 0 0 | + | 22 28 1 0 0 0 0 |
| − | 23 29 1 0 0 0 0 | + | 23 29 1 0 0 0 0 |
| − | 26 30 1 0 0 0 0 | + | 26 30 1 0 0 0 0 |
| − | 30 31 1 0 0 0 0 | + | 30 31 1 0 0 0 0 |
| − | 29 32 1 0 0 0 0 | + | 29 32 1 0 0 0 0 |
| − | 32 33 2 0 0 0 0 | + | 32 33 2 0 0 0 0 |
| − | 32 34 1 0 0 0 0 | + | 32 34 1 0 0 0 0 |
| − | 15 35 1 0 0 0 0 | + | 15 35 1 0 0 0 0 |
| − | 36 37 2 0 0 0 0 | + | 36 37 2 0 0 0 0 |
| − | 37 38 1 0 0 0 0 | + | 37 38 1 0 0 0 0 |
| − | 38 39 2 0 0 0 0 | + | 38 39 2 0 0 0 0 |
| − | 39 40 1 0 0 0 0 | + | 39 40 1 0 0 0 0 |
| − | 40 41 2 0 0 0 0 | + | 40 41 2 0 0 0 0 |
| − | 41 36 1 0 0 0 0 | + | 41 36 1 0 0 0 0 |
| − | 39 42 1 0 0 0 0 | + | 39 42 1 0 0 0 0 |
| − | 38 43 1 0 0 0 0 | + | 38 43 1 0 0 0 0 |
| − | 43 44 1 0 0 0 0 | + | 43 44 1 0 0 0 0 |
| − | 34 45 2 0 0 0 0 | + | 34 45 2 0 0 0 0 |
| − | 45 36 1 0 0 0 0 | + | 45 36 1 0 0 0 0 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL3FACCS0046 | + | ID FL3FACCS0046 |
| − | KNApSAcK_ID C00011122 | + | KNApSAcK_ID C00011122 |
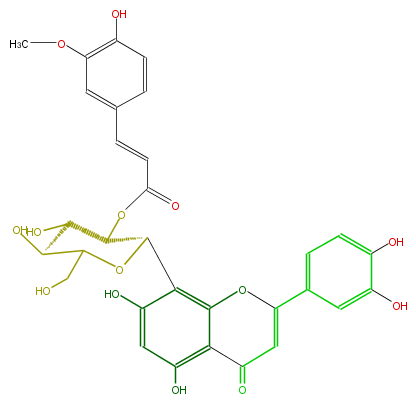
| − | NAME 2''-O-Feruloylorientin | + | NAME 2''-O-Feruloylorientin |
| − | CAS_RN 861691-33-0 | + | CAS_RN 861691-33-0 |
| − | FORMULA C31H28O14 | + | FORMULA C31H28O14 |
| − | EXACTMASS 624.147905604 | + | EXACTMASS 624.147905604 |
| − | AVERAGEMASS 624.5456200000001 | + | AVERAGEMASS 624.5456200000001 |
| − | SMILES C(O1)(CO)C(O)C(O)C(OC(C=Cc(c5)cc(OC)c(c5)O)=O)C(c(c4O)c(c3c(c4)O)OC(=CC3=O)c(c2)cc(c(O)c2)O)1 | + | SMILES C(O1)(CO)C(O)C(O)C(OC(C=Cc(c5)cc(OC)c(c5)O)=O)C(c(c4O)c(c3c(c4)O)OC(=CC3=O)c(c2)cc(c(O)c2)O)1 |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
45 49 0 0 0 0 0 0 0 0999 V2000
-0.6182 1.7612 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0484 2.1460 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0484 2.9157 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.6182 3.3005 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.2847 2.9157 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.2847 2.1460 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7150 1.7612 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.3815 2.1460 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.3815 2.9157 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7150 3.3005 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0199 1.7774 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0199 1.0468 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.6527 0.6814 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.2855 1.0468 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.2855 1.7774 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.6527 2.1428 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.7200 0.7959 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.7150 3.7581 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.6182 3.7603 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.8125 1.8413 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.2941 1.0682 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7543 0.4443 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.0100 0.8001 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.1882 0.7269 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.7279 1.3509 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.4724 0.9951 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.7960 0.5663 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.3688 0.6089 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.6943 0.2533 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.7953 1.5544 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.1956 1.7855 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.1920 -0.2489 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.6245 0.0787 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.1920 -0.8293 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.7960 2.0722 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.8156 -1.8621 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.4012 -2.2002 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.4012 -2.8764 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.8156 -3.2145 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.2299 -2.8764 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.2299 -2.2002 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.8156 -3.7603 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.8942 -3.1610 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.6345 -3.1610 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.8019 -1.1815 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1 2 2 0 0 0 0
2 3 1 0 0 0 0
3 4 2 0 0 0 0
4 5 1 0 0 0 0
5 6 2 0 0 0 0
6 1 1 0 0 0 0
2 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 3 1 0 0 0 0
8 11 1 0 0 0 0
11 12 2 0 0 0 0
12 13 1 0 0 0 0
13 14 2 0 0 0 0
14 15 1 0 0 0 0
15 16 2 0 0 0 0
16 11 1 0 0 0 0
14 17 1 0 0 0 0
10 18 2 0 0 0 0
4 19 1 0 0 0 0
6 20 1 0 0 0 0
21 22 1 1 0 0 0
22 23 1 1 0 0 0
24 23 1 1 0 0 0
24 25 1 0 0 0 0
25 26 1 0 0 0 0
26 21 1 0 0 0 0
1 24 1 0 0 0 0
21 27 1 0 0 0 0
22 28 1 0 0 0 0
23 29 1 0 0 0 0
26 30 1 0 0 0 0
30 31 1 0 0 0 0
29 32 1 0 0 0 0
32 33 2 0 0 0 0
32 34 1 0 0 0 0
15 35 1 0 0 0 0
36 37 2 0 0 0 0
37 38 1 0 0 0 0
38 39 2 0 0 0 0
39 40 1 0 0 0 0
40 41 2 0 0 0 0
41 36 1 0 0 0 0
39 42 1 0 0 0 0
38 43 1 0 0 0 0
43 44 1 0 0 0 0
34 45 2 0 0 0 0
45 36 1 0 0 0 0
S SKP 8
ID FL3FACCS0046
KNApSAcK_ID C00011122
NAME 2''-O-Feruloylorientin
CAS_RN 861691-33-0
FORMULA C31H28O14
EXACTMASS 624.147905604
AVERAGEMASS 624.5456200000001
SMILES C(O1)(CO)C(O)C(O)C(OC(C=Cc(c5)cc(OC)c(c5)O)=O)C(c(c4O)c(c3c(c4)O)OC(=CC3=O)c(c2)cc(c(O)c2)O)1
M END