Mol:FL3FAACS0063
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 53 58 0 0 0 0 0 0 0 0999 V2000 | + | 53 58 0 0 0 0 0 0 0 0999 V2000 |
| − | -0.2608 -1.0868 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.2608 -1.0868 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.2608 -1.8978 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.2608 -1.8978 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.4415 -2.3033 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.4415 -2.3033 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1438 -1.8978 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1438 -1.8978 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1438 -1.0868 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1438 -1.0868 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.4415 -0.6813 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.4415 -0.6813 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8461 -2.3033 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8461 -2.3033 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.5484 -1.8978 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5484 -1.8978 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.5484 -1.0868 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5484 -1.0868 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8461 -0.6813 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8461 -0.6813 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8461 -2.9355 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8461 -2.9355 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.9628 -0.6815 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.9628 -0.6815 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6895 2.2745 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6895 2.2745 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0750 1.6953 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0750 1.6953 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.2494 0.8613 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.2494 0.8613 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.2581 0.0565 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.2581 0.0565 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8430 0.6414 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8430 0.6414 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.4578 1.3710 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.4578 1.3710 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.3047 2.9409 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.3047 2.9409 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.0750 2.4752 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0750 2.4752 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.2023 0.1566 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.2023 0.1566 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.4415 -3.1139 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.4415 -3.1139 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.3286 -0.6554 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.3286 -0.6554 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.0687 -1.0827 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.0687 -1.0827 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.8087 -0.6554 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.8087 -0.6554 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.8087 0.1991 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.8087 0.1991 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.0687 0.6264 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.0687 0.6264 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.3286 0.1991 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.3286 0.1991 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.5484 0.6261 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.5484 0.6261 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7253 -1.8846 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7253 -1.8846 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.2568 -2.5032 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2568 -2.5032 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.5820 -2.2407 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.5820 -2.2407 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.9309 -2.2336 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.9309 -2.2336 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4040 -1.7605 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4040 -1.7605 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.9943 -2.0721 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.9943 -2.0721 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.5519 -1.2375 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.5519 -1.2375 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.9873 -2.5387 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.9873 -2.5387 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.1962 -2.6266 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.1962 -2.6266 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0628 0.6534 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0628 0.6534 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.9788 0.6534 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.9788 0.6534 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.8677 1.3814 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.8677 1.3814 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.0725 1.3304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.0725 1.3304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.4853 2.0455 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.4853 2.0455 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.3110 2.0455 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.3110 2.0455 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.7239 1.3304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.7239 1.3304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.3110 0.6154 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.3110 0.6154 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.4853 0.6154 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.4853 0.6154 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.5484 1.3304 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.5484 1.3304 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.4717 1.3256 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.4717 1.3256 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.5568 3.1139 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.5568 3.1139 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.6548 2.5931 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.6548 2.5931 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8882 1.8014 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8882 1.8014 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.7901 1.2807 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.7901 1.2807 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 1 12 1 0 0 0 0 | + | 1 12 1 0 0 0 0 |
| − | 13 14 1 1 0 0 0 | + | 13 14 1 1 0 0 0 |
| − | 14 15 1 1 0 0 0 | + | 14 15 1 1 0 0 0 |
| − | 16 15 1 1 0 0 0 | + | 16 15 1 1 0 0 0 |
| − | 16 17 1 0 0 0 0 | + | 16 17 1 0 0 0 0 |
| − | 17 18 1 0 0 0 0 | + | 17 18 1 0 0 0 0 |
| − | 18 13 1 0 0 0 0 | + | 18 13 1 0 0 0 0 |
| − | 13 19 1 0 0 0 0 | + | 13 19 1 0 0 0 0 |
| − | 14 20 1 0 0 0 0 | + | 14 20 1 0 0 0 0 |
| − | 15 21 1 0 0 0 0 | + | 15 21 1 0 0 0 0 |
| − | 6 16 1 0 0 0 0 | + | 6 16 1 0 0 0 0 |
| − | 3 22 1 0 0 0 0 | + | 3 22 1 0 0 0 0 |
| − | 23 24 2 0 0 0 0 | + | 23 24 2 0 0 0 0 |
| − | 24 25 1 0 0 0 0 | + | 24 25 1 0 0 0 0 |
| − | 25 26 2 0 0 0 0 | + | 25 26 2 0 0 0 0 |
| − | 26 27 1 0 0 0 0 | + | 26 27 1 0 0 0 0 |
| − | 27 28 2 0 0 0 0 | + | 27 28 2 0 0 0 0 |
| − | 28 23 1 0 0 0 0 | + | 28 23 1 0 0 0 0 |
| − | 23 9 1 0 0 0 0 | + | 23 9 1 0 0 0 0 |
| − | 26 29 1 0 0 0 0 | + | 26 29 1 0 0 0 0 |
| − | 30 31 1 1 0 0 0 | + | 30 31 1 1 0 0 0 |
| − | 31 32 1 1 0 0 0 | + | 31 32 1 1 0 0 0 |
| − | 33 32 1 1 0 0 0 | + | 33 32 1 1 0 0 0 |
| − | 33 34 1 0 0 0 0 | + | 33 34 1 0 0 0 0 |
| − | 34 35 1 0 0 0 0 | + | 34 35 1 0 0 0 0 |
| − | 35 30 1 0 0 0 0 | + | 35 30 1 0 0 0 0 |
| − | 30 36 1 0 0 0 0 | + | 30 36 1 0 0 0 0 |
| − | 31 37 1 0 0 0 0 | + | 31 37 1 0 0 0 0 |
| − | 32 38 1 0 0 0 0 | + | 32 38 1 0 0 0 0 |
| − | 33 2 1 0 0 0 0 | + | 33 2 1 0 0 0 0 |
| − | 21 39 1 0 0 0 0 | + | 21 39 1 0 0 0 0 |
| − | 39 40 1 0 0 0 0 | + | 39 40 1 0 0 0 0 |
| − | 39 41 2 0 0 0 0 | + | 39 41 2 0 0 0 0 |
| − | 42 43 2 0 0 0 0 | + | 42 43 2 0 0 0 0 |
| − | 43 44 1 0 0 0 0 | + | 43 44 1 0 0 0 0 |
| − | 44 45 2 0 0 0 0 | + | 44 45 2 0 0 0 0 |
| − | 45 46 1 0 0 0 0 | + | 45 46 1 0 0 0 0 |
| − | 46 47 2 0 0 0 0 | + | 46 47 2 0 0 0 0 |
| − | 47 42 1 0 0 0 0 | + | 47 42 1 0 0 0 0 |
| − | 45 48 1 0 0 0 0 | + | 45 48 1 0 0 0 0 |
| − | 42 49 1 0 0 0 0 | + | 42 49 1 0 0 0 0 |
| − | 49 40 2 0 0 0 0 | + | 49 40 2 0 0 0 0 |
| − | 50 51 1 0 0 0 0 | + | 50 51 1 0 0 0 0 |
| − | 44 50 1 0 0 0 0 | + | 44 50 1 0 0 0 0 |
| − | 52 53 1 0 0 0 0 | + | 52 53 1 0 0 0 0 |
| − | 18 52 1 0 0 0 0 | + | 18 52 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 50 51 | + | M SAL 1 2 50 51 |
| − | M SBL 1 1 56 | + | M SBL 1 1 56 |
| − | M SMT 1 ^OCH3 | + | M SMT 1 ^OCH3 |
| − | M SBV 1 56 0.2458 -1.0684 | + | M SBV 1 56 0.2458 -1.0684 |
| − | M STY 1 2 SUP | + | M STY 1 2 SUP |
| − | M SLB 1 2 2 | + | M SLB 1 2 2 |
| − | M SAL 2 2 52 53 | + | M SAL 2 2 52 53 |
| − | M SBL 2 1 58 | + | M SBL 2 1 58 |
| − | M SMT 2 CH2OH | + | M SMT 2 CH2OH |
| − | M SBV 2 58 -0.4304 -0.4304 | + | M SBV 2 58 -0.4304 -0.4304 |
| − | S SKP 5 | + | S SKP 5 |
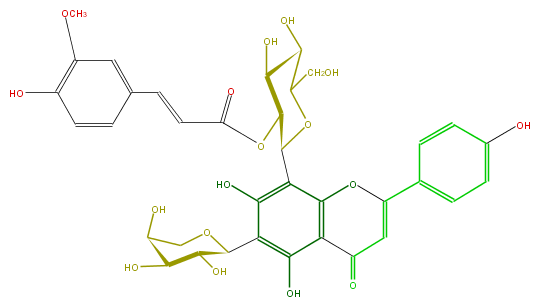
| − | ID FL3FAACS0063 | + | ID FL3FAACS0063 |
| − | FORMULA C36H36O17 | + | FORMULA C36H36O17 |
| − | EXACTMASS 740.1952497259999 | + | EXACTMASS 740.1952497259999 |
| − | AVERAGEMASS 740.66084 | + | AVERAGEMASS 740.66084 |
| − | SMILES C(=CC(OC(C(O)6)C(OC(CO)C(O)6)c(c(O)4)c(c(c(O)c4C(O5)C(C(C(O)C5)O)O)3)OC(=CC3=O)c(c2)ccc(c2)O)=O)c(c1)cc(c(O)c1)OC | + | SMILES C(=CC(OC(C(O)6)C(OC(CO)C(O)6)c(c(O)4)c(c(c(O)c4C(O5)C(C(C(O)C5)O)O)3)OC(=CC3=O)c(c2)ccc(c2)O)=O)c(c1)cc(c(O)c1)OC |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
53 58 0 0 0 0 0 0 0 0999 V2000
-0.2608 -1.0868 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.2608 -1.8978 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.4415 -2.3033 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1438 -1.8978 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1438 -1.0868 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.4415 -0.6813 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8461 -2.3033 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.5484 -1.8978 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.5484 -1.0868 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8461 -0.6813 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.8461 -2.9355 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.9628 -0.6815 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.6895 2.2745 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.0750 1.6953 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.2494 0.8613 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.2581 0.0565 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.8430 0.6414 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.4578 1.3710 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.3047 2.9409 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.0750 2.4752 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.2023 0.1566 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.4415 -3.1139 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.3286 -0.6554 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.0687 -1.0827 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.8087 -0.6554 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.8087 0.1991 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.0687 0.6264 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.3286 0.1991 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.5484 0.6261 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.7253 -1.8846 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.2568 -2.5032 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.5820 -2.2407 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.9309 -2.2336 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.4040 -1.7605 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.9943 -2.0721 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.5519 -1.2375 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.9873 -2.5387 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.1962 -2.6266 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.0628 0.6534 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.9788 0.6534 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.8677 1.3814 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.0725 1.3304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.4853 2.0455 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.3110 2.0455 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.7239 1.3304 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.3110 0.6154 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.4853 0.6154 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.5484 1.3304 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.4717 1.3256 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.5568 3.1139 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.6548 2.5931 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.8882 1.8014 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.7901 1.2807 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
1 12 1 0 0 0 0
13 14 1 1 0 0 0
14 15 1 1 0 0 0
16 15 1 1 0 0 0
16 17 1 0 0 0 0
17 18 1 0 0 0 0
18 13 1 0 0 0 0
13 19 1 0 0 0 0
14 20 1 0 0 0 0
15 21 1 0 0 0 0
6 16 1 0 0 0 0
3 22 1 0 0 0 0
23 24 2 0 0 0 0
24 25 1 0 0 0 0
25 26 2 0 0 0 0
26 27 1 0 0 0 0
27 28 2 0 0 0 0
28 23 1 0 0 0 0
23 9 1 0 0 0 0
26 29 1 0 0 0 0
30 31 1 1 0 0 0
31 32 1 1 0 0 0
33 32 1 1 0 0 0
33 34 1 0 0 0 0
34 35 1 0 0 0 0
35 30 1 0 0 0 0
30 36 1 0 0 0 0
31 37 1 0 0 0 0
32 38 1 0 0 0 0
33 2 1 0 0 0 0
21 39 1 0 0 0 0
39 40 1 0 0 0 0
39 41 2 0 0 0 0
42 43 2 0 0 0 0
43 44 1 0 0 0 0
44 45 2 0 0 0 0
45 46 1 0 0 0 0
46 47 2 0 0 0 0
47 42 1 0 0 0 0
45 48 1 0 0 0 0
42 49 1 0 0 0 0
49 40 2 0 0 0 0
50 51 1 0 0 0 0
44 50 1 0 0 0 0
52 53 1 0 0 0 0
18 52 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 50 51
M SBL 1 1 56
M SMT 1 ^OCH3
M SBV 1 56 0.2458 -1.0684
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 52 53
M SBL 2 1 58
M SMT 2 CH2OH
M SBV 2 58 -0.4304 -0.4304
S SKP 5
ID FL3FAACS0063
FORMULA C36H36O17
EXACTMASS 740.1952497259999
AVERAGEMASS 740.66084
SMILES C(=CC(OC(C(O)6)C(OC(CO)C(O)6)c(c(O)4)c(c(c(O)c4C(O5)C(C(C(O)C5)O)O)3)OC(=CC3=O)c(c2)ccc(c2)O)=O)c(c1)cc(c(O)c1)OC
M END