Mol:FL1DA9NC0005
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 36 39 0 0 0 0 0 0 0 0999 V2000 | + | 36 39 0 0 0 0 0 0 0 0999 V2000 |
| − | -0.5308 -0.3231 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5308 -0.3231 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5308 -0.9628 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5308 -0.9628 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0232 -1.2827 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0232 -1.2827 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5772 -0.9628 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5772 -0.9628 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5772 -0.3231 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5772 -0.3231 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0232 -0.0033 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0232 -0.0033 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1309 -1.2825 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1309 -1.2825 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.6834 -0.9635 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.6834 -0.9635 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2348 -1.2818 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2348 -1.2818 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.7849 -0.9642 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.7849 -0.9642 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.3233 -1.2750 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.3233 -1.2750 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.8617 -0.9642 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.8617 -0.9642 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.8617 -0.3425 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.8617 -0.3425 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.3233 -0.0317 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.3233 -0.0317 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.7849 -0.3425 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.7849 -0.3425 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1309 -1.9205 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1309 -1.9205 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0232 -1.9221 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0232 -1.9221 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0845 -0.0034 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0845 -0.0034 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0845 -1.2825 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0845 -1.2825 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6382 -0.9628 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6382 -0.9628 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6382 -0.3023 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6382 -0.3023 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.2102 0.0279 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2102 0.0279 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7822 -0.3023 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7822 -0.3023 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7822 -0.9628 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7822 -0.9628 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.2102 -1.2931 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2102 -1.2931 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.2102 -1.9531 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2102 -1.9531 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.2102 0.6879 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2102 0.6879 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7818 1.0179 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7818 1.0179 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7818 1.6414 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7818 1.6414 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.3217 1.9531 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.3217 1.9531 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.8617 1.6414 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.8617 1.6414 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.8617 1.0179 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.8617 1.0179 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.3217 0.7062 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.3217 0.7062 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.2424 1.9528 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.2424 1.9528 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1309 -0.0034 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1309 -0.0034 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8454 -0.4159 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8454 -0.4159 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 2 0 0 0 0 | + | 1 2 2 0 0 0 0 |
| − | 2 3 1 0 0 0 0 | + | 2 3 1 0 0 0 0 |
| − | 3 4 2 0 0 0 0 | + | 3 4 2 0 0 0 0 |
| − | 4 5 1 0 0 0 0 | + | 4 5 1 0 0 0 0 |
| − | 5 6 2 0 0 0 0 | + | 5 6 2 0 0 0 0 |
| − | 6 1 1 0 0 0 0 | + | 6 1 1 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 1 0 0 0 0 | + | 8 9 1 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 11 2 0 0 0 0 | + | 10 11 2 0 0 0 0 |
| − | 11 12 1 0 0 0 0 | + | 11 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 10 1 0 0 0 0 | + | 15 10 1 0 0 0 0 |
| − | 7 16 2 0 0 0 0 | + | 7 16 2 0 0 0 0 |
| − | 3 17 1 0 0 0 0 | + | 3 17 1 0 0 0 0 |
| − | 1 18 1 0 0 0 0 | + | 1 18 1 0 0 0 0 |
| − | 2 19 1 0 0 0 0 | + | 2 19 1 0 0 0 0 |
| − | 19 20 1 0 0 0 0 | + | 19 20 1 0 0 0 0 |
| − | 20 21 2 0 0 0 0 | + | 20 21 2 0 0 0 0 |
| − | 21 22 1 0 0 0 0 | + | 21 22 1 0 0 0 0 |
| − | 22 23 2 0 0 0 0 | + | 22 23 2 0 0 0 0 |
| − | 23 24 1 0 0 0 0 | + | 23 24 1 0 0 0 0 |
| − | 24 25 2 0 0 0 0 | + | 24 25 2 0 0 0 0 |
| − | 25 20 1 0 0 0 0 | + | 25 20 1 0 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 22 27 1 0 0 0 0 | + | 22 27 1 0 0 0 0 |
| − | 27 28 1 0 0 0 0 | + | 27 28 1 0 0 0 0 |
| − | 28 29 2 0 0 0 0 | + | 28 29 2 0 0 0 0 |
| − | 29 30 1 0 0 0 0 | + | 29 30 1 0 0 0 0 |
| − | 30 31 2 0 0 0 0 | + | 30 31 2 0 0 0 0 |
| − | 31 32 1 0 0 0 0 | + | 31 32 1 0 0 0 0 |
| − | 32 33 2 0 0 0 0 | + | 32 33 2 0 0 0 0 |
| − | 33 28 1 0 0 0 0 | + | 33 28 1 0 0 0 0 |
| − | 29 34 1 0 0 0 0 | + | 29 34 1 0 0 0 0 |
| − | 5 35 1 0 0 0 0 | + | 5 35 1 0 0 0 0 |
| − | 35 36 1 0 0 0 0 | + | 35 36 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 35 36 | + | M SAL 1 2 35 36 |
| − | M SBL 1 1 38 | + | M SBL 1 1 38 |
| − | M SMT 1 OCH3 | + | M SMT 1 OCH3 |
| − | M SBV 1 38 -4.2836 2.7809 | + | M SBV 1 38 -4.2836 2.7809 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL1DA9NC0005 | + | ID FL1DA9NC0005 |
| − | KNApSAcK_ID C00007964 | + | KNApSAcK_ID C00007964 |
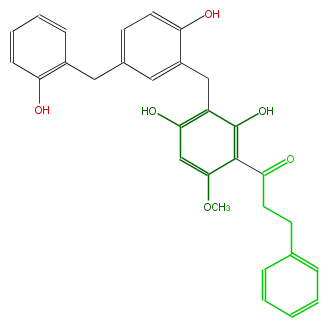
| − | NAME Angoluvarin | + | NAME Angoluvarin |
| − | CAS_RN 110874-65-2 | + | CAS_RN 110874-65-2 |
| − | FORMULA C30H28O6 | + | FORMULA C30H28O6 |
| − | EXACTMASS 484.188588628 | + | EXACTMASS 484.188588628 |
| − | AVERAGEMASS 484.53972000000005 | + | AVERAGEMASS 484.53972000000005 |
| − | SMILES c(c4)(cccc4)CCC(=O)c(c(O)1)c(cc(O)c1Cc(c2O)cc(Cc(c3O)cccc3)cc2)OC | + | SMILES c(c4)(cccc4)CCC(=O)c(c(O)1)c(cc(O)c1Cc(c2O)cc(Cc(c3O)cccc3)cc2)OC |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
36 39 0 0 0 0 0 0 0 0999 V2000
-0.5308 -0.3231 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5308 -0.9628 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0232 -1.2827 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5772 -0.9628 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5772 -0.3231 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0232 -0.0033 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1309 -1.2825 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.6834 -0.9635 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.2348 -1.2818 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.7849 -0.9642 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.3233 -1.2750 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.8617 -0.9642 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.8617 -0.3425 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.3233 -0.0317 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.7849 -0.3425 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1309 -1.9205 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.0232 -1.9221 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.0845 -0.0034 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.0845 -1.2825 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6382 -0.9628 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6382 -0.3023 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.2102 0.0279 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7822 -0.3023 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7822 -0.9628 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.2102 -1.2931 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.2102 -1.9531 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.2102 0.6879 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7818 1.0179 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7818 1.6414 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.3217 1.9531 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.8617 1.6414 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.8617 1.0179 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.3217 0.7062 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.2424 1.9528 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.1309 -0.0034 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.8454 -0.4159 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1 2 2 0 0 0 0
2 3 1 0 0 0 0
3 4 2 0 0 0 0
4 5 1 0 0 0 0
5 6 2 0 0 0 0
6 1 1 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 1 0 0 0 0
9 10 1 0 0 0 0
10 11 2 0 0 0 0
11 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 10 1 0 0 0 0
7 16 2 0 0 0 0
3 17 1 0 0 0 0
1 18 1 0 0 0 0
2 19 1 0 0 0 0
19 20 1 0 0 0 0
20 21 2 0 0 0 0
21 22 1 0 0 0 0
22 23 2 0 0 0 0
23 24 1 0 0 0 0
24 25 2 0 0 0 0
25 20 1 0 0 0 0
25 26 1 0 0 0 0
22 27 1 0 0 0 0
27 28 1 0 0 0 0
28 29 2 0 0 0 0
29 30 1 0 0 0 0
30 31 2 0 0 0 0
31 32 1 0 0 0 0
32 33 2 0 0 0 0
33 28 1 0 0 0 0
29 34 1 0 0 0 0
5 35 1 0 0 0 0
35 36 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 35 36
M SBL 1 1 38
M SMT 1 OCH3
M SBV 1 38 -4.2836 2.7809
S SKP 8
ID FL1DA9NC0005
KNApSAcK_ID C00007964
NAME Angoluvarin
CAS_RN 110874-65-2
FORMULA C30H28O6
EXACTMASS 484.188588628
AVERAGEMASS 484.53972000000005
SMILES c(c4)(cccc4)CCC(=O)c(c(O)1)c(cc(O)c1Cc(c2O)cc(Cc(c3O)cccc3)cc2)OC
M END