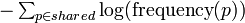Template:MassBank/Matrix
From Metabolomics.JP
(Difference between revisions)
m |
|||
| (194 intermediate revisions by one user not shown) | |||
| Line 1: | Line 1: | ||
| − | < | + | {{MassBank/Header}} |
| − | {| class ="wikitable" | + | {{#def:MASSBANKID|{{#substring:{{PAGENAME}}|0|{{#expr:{{#length:{{PAGENAME}}}}-1}}}}}} |
| + | {{#def:dataline|{{#SearchLine:{{PAGENAME}}|Index|MassBank/Molecules}}}} | ||
| + | <!---化合物組成式の正規表現--------->{{#def:FORMULA_PAT|(C?[1-9]?[0-9]?)(H?[1-9]?[0-9]?)(C?l?[2-9]?)(F?[2-9]?)(I?[2-9]?)(N?[1-9]?[0-9]?)(O?[1-9]?[0-9]?)(P?[2-9]?)(S?[2-9]?)}} | ||
| + | <!---化合物組成式に出てくる文字列--->{{#def:FORMULA_CHAR|CHFINOPSl0-9%*}} | ||
| + | <!---化合物組成式に使われる原子----->{{#def:ATOM|{"C", "H", "Cl", "F", "I", "N", "O", "P", "S"} }} | ||
| + | <!---各原子の質量(上のリスト)------->{{#def:MASS|{12, 1, 35, 19, 127, 14, 16, 31, 32} }} | ||
| + | <!---化合物の名前------------------->{{#def:NAME|{{#nth:{{#var:dataline}}|6|&&}}}} | ||
| + | |||
| + | <span style="float:right;"> | ||
| + | [[Image:{{#var:MASSBANKID}}.png|150px]]<br/> | ||
| + | <big>{{#var:NAME}}</big><br/> | ||
| + | </span> | ||
| + | |||
| + | <div> | ||
| + | {| class="wikitable nowraplinks" border="1" cellpadding="2" cellspacing="1" style="float: right; clear: none; margin: 1em 1em 1em 1em" | ||
| + | |- | ||
| + | ! colspan="2" | IDs and Links | ||
| + | |- class | ||
| + | | style="width: 35%;" | [http://massbank.jp/ MassBank] | ||
| + | | style="width: 65%;" | [http://www.massbank.jp/jsp/FwdRecord.jsp?id={{#var:MASSBANKID}} {{#var:NAME}}] | ||
| + | |- class{{#nth:{{#var:dataline}}|3|&&}}="hiddenStructure" | ||
| + | | style="width: 35%;" | [https://scifinder.cas.org/ CAS] | ||
| + | | style="width: 65%;" | {{#nth:{{#var:dataline}}|3|&&}} | ||
| + | |- class | ||
| + | | style="width: 35%;" | Keio ID | ||
| + | | style="width: 65%;" | {{#nth:{{#var:dataline}}|4|&&}} | ||
| + | |} | ||
| + | |||
| + | ==Top 10 Similar Molecules of {{#var:NAME}}== | ||
| + | {| class="wikitable collapsible collapsed" | ||
| + | ! Ranking | ||
| + | ! About scoring | ||
| + | |- | ||
| + | | | ||
| + | {{#repeat:MassBank/Matrix/RankingLink|1| | ||
| + | {{#lua: | ||
| + | function intersection(x, y) | ||
| + | local z = {} | ||
| + | local i = 1 | ||
| + | local j = 1 | ||
| + | repeat | ||
| + | if (x[i] == y[j]) | ||
| + | then table.insert(z, x[i]) i = i + 1 j = j + 1 end | ||
| + | if (i > table.getn(x) or j > table.getn(y)) | ||
| + | then break end | ||
| + | if (x[i] > y[j]) then j = j + 1 end | ||
| + | if (j > table.getn(y)) then break end | ||
| + | if (x[i] < y[j]) then i = i + 1 end | ||
| + | if (i > table.getn(x)) then break end | ||
| + | until false | ||
| + | return z | ||
| + | end | ||
| + | |||
| + | Name = {} | ||
| + | Ions = {} | ||
| + | Freq = {} | ||
| + | total = 0 | ||
| + | ---Register Ions--- | ||
| + | ---全ページのイオンヘッダーにおける頻度を登録。最初の要素はページ名--- | ||
| + | for id, frag in stdin:gmatch("&&([%a%d]+)&&([%S]+)") do | ||
| + | table.insert(Name, id) | ||
| + | T = {} | ||
| + | for f in string.gmatch(frag,"([%a%d]+)&&") do | ||
| + | table.insert(T, f) | ||
| + | if (Freq[f]) then Freq[f] = Freq[f] + 1 else Freq[f] = 1 total = total + 1 end | ||
| + | end | ||
| + | table.sort(T) | ||
| + | table.insert(Ions, T) | ||
| + | end | ||
| + | ---Compute Similarity--- | ||
| + | Rank = {} | ||
| + | Score = {} | ||
| + | for i=2, table.getn(Ions) do | ||
| + | R = intersection(Ions[i], Ions[1]) | ||
| + | if (table.getn(R) > 0) then | ||
| + | inf = 0 | ||
| + | for _,v in pairs(R) do | ||
| + | inf = inf - math.log(Freq[v]/total) | ||
| + | end | ||
| + | inf = math.floor(inf*100) + i * 0.0001 | ||
| + | table.insert(Score, inf) | ||
| + | Rank[inf] = i | ||
| + | end | ||
| + | end | ||
| + | table.sort(Score) | ||
| + | ---Output Similarity--- | ||
| + | for i=table.getn(Score), math.max(table.getn(Score)-9,1), -1 do | ||
| + | print(Name[ Rank[Score[i]] ] .. " Score:" ..math.floor(Score[i])/100) | ||
| + | end | ||
| + | | | ||
| + | &&{{PAGENAME}}&&{{#car:{{{data}}}}} | ||
| + | {{#SearchLine:^&&_%&&|MassBank}} | ||
| + | }} | ||
| + | }} | ||
| + | |valign="top"| | ||
| + | The similarity score between two molecules is defined as the sum of Shannon entropy of ionic formulas shared by the molecules: | ||
| + | <math>\textstyle - \sum_{p\in shared} \log(\mbox{frequency}(p))</math> | ||
| + | <br/> | ||
| + | 代謝物間の類似度スコアは、それらのスペクトルで共有される(アノテートされた)イオン式のシャノン情報量の総和です。 | ||
| + | |} | ||
| + | |||
| + | |||
| + | =Links= | ||
| + | {| class="wikitable collapsible collapsed" | ||
| + | ! Page list for each fragment | ||
| + | |- | ||
| + | | | ||
| + | {{#repeat:MassBank/SearchFgmt|1| | ||
| + | {{#lua: | ||
| + | ---find and print Ruler--- | ||
| + | ---ヘッダー部分を分解するだけ--- | ||
| + | for line in stdin:gmatch("[%S ,]+") do | ||
| + | if (string.find(line, "^&&[&%a%d]+&&$") ~= nil) then | ||
| + | print(line) return | ||
| + | end | ||
| + | end | ||
| + | |{{{data|CHClFINOPS}}} | ||
| + | }}|&&}} | ||
| + | |} | ||
| + | |||
| + | ==Annotations== | ||
| + | {{#ifeq:{{NAMESPACE}}|Ojima| | ||
| + | <div style="float:right"> | ||
| + | {{#ifexists:Image:Fragmentation:{{PAGENAME}}.png|[[Image:Fragmentation:{{PAGENAME}}.png]]}} | ||
| + | </div> | ||
| + | }} | ||
{{#replace: | {{#replace: | ||
{{#lua: | {{#lua: | ||
| − | function | + | ---print comments--- |
| − | + | FORMULA_PAT = "{{#var:FORMULA_PAT}}" | |
| − | + | FORMULA_CHAR = "{{#var:FORMULA_CHAR}}" | |
| − | + | ATOM = {{#var:ATOM}} | |
| − | + | MASS = {{#var:MASS}} | |
| − | + | ||
| + | function toFormula(t) | ||
| + | for i,v in pairs(t) do | ||
| + | if (v == "") | ||
| + | then t[i] = 0 | ||
| + | else if (v == ATOM[i]) | ||
| + | then t[i] = 1 | ||
| + | else t[i]=tonumber(string.sub(v,1+string.len(ATOM[i]))) | ||
| + | end | ||
| + | end | ||
end | end | ||
| + | return t | ||
end | end | ||
function mass(str) | function mass(str) | ||
| − | local | + | local t = toFormula({string.match(str,FORMULA_PAT)}) |
| − | + | ret = 0; | |
| − | + | for i,v in pairs(t) do | |
| − | + | ret = ret + t[i] * MASS[i] | |
| − | + | end | |
| − | + | return ret; | |
| − | + | end | |
| − | + | ||
| − | + | ret = "" | |
| + | comment = ""; | ||
| + | head = ""; | ||
| + | tail = ""; | ||
| + | for line in stdin:gmatch("[%S ,]+") do | ||
| + | repeat | ||
| + | if (string.find(line, "^&&[&%a%d]+&&$") ~= nil) then break end | ||
| + | local h, t = string.match(line, "^&&(["..FORMULA_CHAR.."]+) *: *(["..FORMULA_CHAR.." ]+)$"); | ||
| + | if (h ~= nil and t ~= nil) then | ||
| + | if (comment ~= "") then | ||
| + | ret = ret .."~-\n~~"..head.." ('''"..mass(head).."''')".." ~~ "..tail.." ('''"..mass(tail).."''')".."\n~~ "..comment.."\n" | ||
| + | comment = "" | ||
| + | end | ||
| + | head = h; tail = t | ||
| + | break | ||
| + | end | ||
| + | ---comment lines--- | ||
| + | comment = comment .. line.."\n" | ||
| + | until true | ||
| + | end | ||
| + | ---process the last comment (head=precursor, tail=product)--- | ||
| + | if (comment ~= "") then | ||
| + | ret = ret .. "~-\n~~"..head.." ('''"..mass(head).."''')".." ~~ "..tail.." ('''"..mass(tail).."''')".."\n~~ "..comment.."\n" | ||
| + | end | ||
| + | if (ret ~= "") then | ||
| + | ret = '{~ class="wikitable sortable"\n!Precursor~~Product~~Comments\n'.. ret ..'~}' | ||
| + | print(ret) | ||
| + | end | ||
| + | |{{{data|CHClFINOPS}}} | ||
| + | }} | ||
| + | |~|{{#bar:}}}} | ||
| + | |||
| + | ==Precursor-Product Relationship== | ||
| + | {| class="collapsible collapsed" | ||
| + | ! colspan="2"| About the PP Table (行列表示について) | ||
| + | |- | ||
| + | |The matrix is viewed columnwise. The topmost precursor ion in bold face produces the product ions beneath it. | ||
| + | Each formula in matrix cells corresponds to the neutral loss. Blackout cells indicate products that cannot be derived, and orange cells indicate a structurally plausible link produced by cleaving a single chemical bond (in cases of ring-opening, two bonds). | ||
| + | |行列は列方向に見ます。最上段太字の前駆イオン(precursor ion)が直下の生成イオン群(product ions)になると解釈します。 | ||
| + | 行列要素に書かれている組成式はニュートラルロスです。黒は前駆イオンから生成しえない関係、オレンジは分子構造における共有結合1本の切断(開環の場合は2本)で生じる関係を意味します。 | ||
| + | |} | ||
| + | {{#replace: | ||
| + | {{#lua: | ||
| + | FORMULA_PAT = "{{#var:FORMULA_PAT}}" | ||
| + | FORMULA_CHAR = "{{#var:FORMULA_CHAR}}" | ||
| + | ATOM = {{#var:ATOM}} | ||
| + | MASS = {{#var:MASS}} | ||
| + | |||
| + | function toFormula(t) | ||
| + | for i,v in pairs(t) do | ||
| + | if (v == "") | ||
| + | then t[i] = 0 | ||
| + | else if (v == ATOM[i]) | ||
| + | then t[i] = 1 | ||
| + | else t[i]=tonumber(string.sub(v,1+string.len(ATOM[i]))) | ||
| + | end | ||
| + | end | ||
| + | end | ||
| + | return t | ||
| + | end | ||
| + | |||
| + | function mass(str) | ||
| + | local t = toFormula({string.match(str,FORMULA_PAT)}) | ||
| + | ret = 0; | ||
| + | for i,v in pairs(t) do | ||
| + | ret = ret + t[i] * MASS[i] | ||
| + | end | ||
| + | return ret; | ||
end | end | ||
function diff(str1, str2) | function diff(str1, str2) | ||
---computes str1 - str2. If negative, returns nil.--- | ---computes str1 - str2. If negative, returns nil.--- | ||
| − | local | + | local t1 = toFormula({string.match(str1,FORMULA_PAT)}) |
| − | local | + | local t2 = toFormula({string.match(str2,FORMULA_PAT)}) |
| − | + | for i,_ in pairs(t1) do | |
| − | + | if (t1[i] < t2[i]) then return nil else t1[i] = t1[i]-t2[i] end | |
| − | + | end | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
local ret = ""; | local ret = ""; | ||
| − | + | for i,v in pairs(t1) do | |
| − | + | if (v >= 1) then ret = ret .. ATOM[i] end | |
| − | + | if (v > 1) then ret = ret .. v end | |
| − | + | end | |
| − | + | return ret | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | return ret | + | |
end | end | ||
| − | local ruler; | + | ---Main Program--- |
| − | local fragments = {}; | + | local ruler = nil; ---first line (list of fragments)--- |
| + | local fragments = {}; ---following lines (head : tail style)--- | ||
| + | local focus = {}; ---formulas that should be highlighted--- | ||
local x, y; | local x, y; | ||
| + | ---Read Data--- | ||
for line in stdin:gmatch("[%S ,]+") do | for line in stdin:gmatch("[%S ,]+") do | ||
| − | if (string.find(line, " | + | if (string.find(line, "^&&[&%a%d]+&&$") ~= nil) |
| − | + | then ruler = string.gsub(line, "&&", " ") | |
| − | head, tail = string.match(line, "([ | + | elseif (string.match(line,"^&&")) then |
| − | + | ---register fragments--- | |
| − | + | head, tail = string.match(line, "^&&(["..FORMULA_CHAR.."]+) *: *(["..FORMULA_CHAR.." ]+)$") | |
| − | + | if (string.match(head,"*")) then | |
| + | head = string.gsub(head,"*", "") | ||
| + | focus[head] = head | ||
end | end | ||
| − | fragments[head] = y | + | if (head ~= nil and tail ~= nil) then |
| + | if (fragments[head] == nil) | ||
| + | then y = {} else y = fragments[head] end | ||
| + | for x in string.gmatch(tail,"["..FORMULA_CHAR.."]+") do | ||
| + | z = string.gsub(x,"*", "") | ||
| + | y[x] = z | ||
| + | y[z] = z | ||
| + | end | ||
| + | fragments[head] = y | ||
| + | else | ||
| + | print('<span style="color:red">Ion in '..line..' does not exist.</span>') | ||
| + | end | ||
| + | end | ||
| + | end | ||
| + | ---Check Ruler--- | ||
| + | if (ruler == nil) then | ||
| + | print('<span style="color:red">No "ION INFO" line in the form &&(formula)&&(formula)&&...&&</span>') | ||
| + | ruler = "CHClFIONSP" | ||
| + | end | ||
| + | x = nil | ||
| + | inRuler = {} | ||
| + | for formula in string.gmatch(ruler, "([%S]+)") do | ||
| + | if (x ~= nil and mass(formula) > mass(x)) then | ||
| + | print('<span style="color:red">Illegal ion order (check mass!): '..x..' < '..formula..'</span><br/>') | ||
| + | end | ||
| + | x = formula | ||
| + | inRuler[formula] = true | ||
| + | end | ||
| + | ---Check Fragments--- | ||
| + | for i,v in pairs(fragments) do | ||
| + | ---i is head, v is a list of fragments--- | ||
| + | if (not inRuler[i]) then print('<span style="color:red">Parent ion '..i..' does not exist.</span>') end | ||
| + | for j,w in pairs(v) do | ||
| + | j = string.gsub(j,"*","") | ||
| + | if (not inRuler[w]) then print('<span style="color:red">Fragment ion '..w..' does not exist.</span>') end | ||
end | end | ||
end | end | ||
| + | ---Write Matrix--- | ||
| + | print("<small>") | ||
| + | print('{# class ="wikitable collapsible" style="margin:0 0 0 0"') | ||
local axis = {}; | local axis = {}; | ||
local i = 1; | local i = 1; | ||
for formula in string.gmatch(ruler, "([%S]+)") do | for formula in string.gmatch(ruler, "([%S]+)") do | ||
| − | axis[i] = formula | + | axis[i] = formula |
| − | i = i + 1 | + | i = i + 1 |
end | end | ||
| − | print("#-\n") | + | ---Header--- |
| − | print(" | + | print("#-\n") |
| + | print("! {{PAGENAME}} \n") | ||
for i=1, table.getn(axis) do | for i=1, table.getn(axis) do | ||
| − | print("#style='text-align:right'# " .. mass(axis[i]) .. "<br/>" .. axis[i]) | + | ---If some fragments are focused then highlight the axis--- |
| + | if (focus[axis[i]] ~= nil) then | ||
| + | print("#style='text-align:right;background:red'# '''" .. mass(axis[i]) .. "'''<br/>" .. axis[i]) | ||
| + | else | ||
| + | print("#style='text-align:right'# '''" .. mass(axis[i]) .. "'''<br/>" .. axis[i]) | ||
| + | end | ||
end | end | ||
| + | ---Rows--- | ||
for i=1, table.getn(axis) do | for i=1, table.getn(axis) do | ||
| − | print("#-\n") | + | print("#-\n") |
| − | print("#style='text-align:right'# " .. mass(axis[i]) .. " | + | print("#style='text-align:right'# '''" .. mass(axis[i]) .. "''' " .. axis[i]) |
for j=1, table.getn(axis) do | for j=1, table.getn(axis) do | ||
if (j < i) then | if (j < i) then | ||
s = diff(axis[j],axis[i]); | s = diff(axis[j],axis[i]); | ||
if (s == nil) | if (s == nil) | ||
| − | then print('#style="background-color:gray"# ') | + | then print('#style="background-color:gray"# ') |
else | else | ||
if (fragments[axis[j]] ~= nil and fragments[axis[j]][axis[i]] ~= nil) then | if (fragments[axis[j]] ~= nil and fragments[axis[j]][axis[i]] ~= nil) then | ||
| − | print('#style="background-color:orange"# ' .. s) | + | ---記号の有無により色を変更--- |
| − | else print('## ' .. s) | + | if (fragments[axis[j]][axis[i].."*"] ~= nil) then |
| + | print('#style="background-color:skyblue"# ' .. s) | ||
| + | elseif (fragments[axis[j]][axis[i].."**"] ~= nil) then | ||
| + | print('#style="background-color:orange"# ' .. s) | ||
| + | elseif (fragments[axis[j]][axis[i].."***"] ~= nil) then | ||
| + | print('#style="background-color:beige"# ' .. s) | ||
| + | elseif (fragments[axis[j]][axis[i].."****"] ~= nil) then | ||
| + | print('#style="background-color:coral"# ' .. s) | ||
| + | else | ||
| + | print('#style="background-color:orange"# ' .. s) | ||
| + | end | ||
| + | else print('## ' .. s) end | ||
end | end | ||
| − | else print('#style="background-color:white"# ') | + | else print('#style="background-color:white"# ') |
end | end | ||
end | end | ||
| − | print("\n") | + | print("\n") |
end | end | ||
| + | print("#}") | ||
| + | print("</small>") | ||
| + | ---End of Matrix--- | ||
| | | | ||
| − | {{{data| | + | {{{data|CHCFIlNOPS}}} |
}} | }} | ||
|#|{{#bar:}}}} | |#|{{#bar:}}}} | ||
| − | + | ||
| − | </ | + | <!---- next/prev links at the page bottom----> |
| + | {{#def:KeioID|{{#ifeq:{{#substring:{{PAGENAME}}|0|2}}|KO|{{#substring:{{#nth:{{#SearchLine:{{PAGENAME}}|Index|MassBank/Molecules}}|4|&&}}|0|4}}|{{#substring:{{PAGENAME}}|0|8}}}}}} | ||
| + | {{#repeat:MassBank/PrevNextLink|4| | ||
| + | {{#lua: | ||
| + | first = nil | ||
| + | prev = nil | ||
| + | next = nil | ||
| + | last = nil | ||
| + | hit = nil | ||
| + | for word in stdin:gmatch("(%w+)") do | ||
| + | if (first == nil) then first = word end | ||
| + | last = word | ||
| + | if (word == "{{#var:KeioID}}") then | ||
| + | hit = true | ||
| + | if (tmp ~= nil) then | ||
| + | prev = tmp | ||
| + | else | ||
| + | prev = first | ||
| + | end | ||
| + | elseif (hit) then | ||
| + | next = word | ||
| + | hit = false | ||
| + | end | ||
| + | tmp = word | ||
| + | end | ||
| + | if (next == nil) then next = last end | ||
| + | print(first) | ||
| + | print(prev) | ||
| + | print(next) | ||
| + | print(last) | ||
| + | |{{Persist:MassBank/AllPages}}}} | ||
| + | }} | ||
Latest revision as of 16:27, 7 April 2011
| General Index | Ion Frequency | Prec.-Product | Neutral Loss | Help |
| IDs and Links | |
|---|---|
| MassBank | [1] |
| CAS | |
| Keio ID | |
Contents |
[edit] Top 10 Similar Molecules of
| Ranking | About scoring |
|---|---|
|
The similarity score between two molecules is defined as the sum of Shannon entropy of ionic formulas shared by the molecules:
|
[edit] Links
| Page list for each fragment |
|---|
[edit] Annotations
| Precursor | Product | Comments |
|---|---|---|
| (0) | (0) | CHClFINOPS |
[edit] Precursor-Product Relationship
| About the PP Table (行列表示について) | |
|---|---|
| The matrix is viewed columnwise. The topmost precursor ion in bold face produces the product ions beneath it.
Each formula in matrix cells corresponds to the neutral loss. Blackout cells indicate products that cannot be derived, and orange cells indicate a structurally plausible link produced by cleaving a single chemical bond (in cases of ring-opening, two bonds). |
行列は列方向に見ます。最上段太字の前駆イオン(precursor ion)が直下の生成イオン群(product ions)になると解釈します。
行列要素に書かれている組成式はニュートラルロスです。黒は前駆イオンから生成しえない関係、オレンジは分子構造における共有結合1本の切断(開環の場合は2本)で生じる関係を意味します。 |
No "ION INFO" line in the form &&(formula)&&(formula)&&...&&
| MassBank/Matrix | 210 CHClFIONSP |
|---|---|
| 210 CHClFIONSP |
|