Mol:PR100620
From Metabolomics.JP
(Difference between revisions)
| Line 2: | Line 2: | ||
| − | 39 41 0 0 1 0 0 0 0 0999 V2000 | + | 39 41 0 0 1 0 0 0 0 0999 V2000 |
| − | 32.0842 -27.6135 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 | + | 32.0842 -27.6135 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 |
| − | 33.1497 -25.4069 0.0000 N 0 0 3 0 0 0 0 0 0 0 0 0 | + | 33.1497 -25.4069 0.0000 N 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 30.9722 -26.8043 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 30.9722 -26.8043 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 31.6883 -28.8885 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 31.6883 -28.8885 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 31.9794 -24.6965 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 31.9794 -24.6965 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 34.3492 -24.6965 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 34.3492 -24.6965 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 29.8776 -27.5902 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 29.8776 -27.5902 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 30.3259 -28.8885 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 30.3259 -28.8885 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 32.4918 -29.9948 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 32.4918 -29.9948 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 31.9794 -23.3165 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 | + | 31.9794 -23.3165 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 30.7917 -25.3719 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 30.7917 -25.3719 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 34.3492 -23.3165 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 34.3492 -23.3165 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 28.5850 -27.1826 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 28.5850 -27.1826 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 29.5225 -29.9948 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 29.5225 -29.9948 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 33.1556 -22.6411 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 33.1556 -22.6411 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 28.2939 -25.8493 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 28.2939 -25.8493 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 33.1556 -21.2730 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 33.1556 -21.2730 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 26.9256 -25.8493 0.0000 P 0 0 3 0 0 0 0 0 0 0 0 0 | + | 26.9256 -25.8493 0.0000 P 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 25.5632 -25.8493 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 25.5632 -25.8493 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 26.9256 -24.4811 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 26.9256 -24.4811 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 26.9198 -27.2059 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 26.9198 -27.2059 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 24.2066 -25.8493 0.0000 P 0 0 3 0 0 0 0 0 0 0 0 0 | + | 24.2066 -25.8493 0.0000 P 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 22.8325 -25.8435 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.8325 -25.8435 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 24.2066 -24.4811 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 24.2066 -24.4811 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 24.2066 -27.2059 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 24.2066 -27.2059 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 21.6506 -26.5130 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 21.6506 -26.5130 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 21.6506 -27.8871 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 21.6506 -27.8871 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 20.4629 -25.8435 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 20.4629 -25.8435 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 20.4629 -28.5625 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 | + | 20.4629 -28.5625 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 |
| − | 23.0887 -29.0283 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 | + | 23.0887 -29.0283 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 19.2810 -26.5130 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 19.2810 -26.5130 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 19.2810 -27.8871 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 19.2810 -27.8871 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 20.4629 -29.9249 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 20.4629 -29.9249 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 23.5022 -30.2510 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 23.5022 -30.2510 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 18.1222 -25.8435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 18.1222 -25.8435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 18.1222 -28.5625 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 18.1222 -28.5625 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 24.8471 -30.4198 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 24.8471 -30.4198 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 22.6637 -31.3282 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 22.6637 -31.3282 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 17.0801 -26.7168 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 17.0801 -26.7168 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 1 0 0 | + | 1 2 1 1 0 0 |
| − | 1 3 1 0 0 0 | + | 1 3 1 0 0 0 |
| − | 1 4 1 0 0 0 | + | 1 4 1 0 0 0 |
| − | 2 5 1 0 0 0 | + | 2 5 1 0 0 0 |
| − | 2 6 1 0 0 0 | + | 2 6 1 0 0 0 |
| − | 3 7 1 0 0 0 | + | 3 7 1 0 0 0 |
| − | 4 8 1 0 0 0 | + | 4 8 1 0 0 0 |
| − | 4 9 1 6 0 0 | + | 4 9 1 6 0 0 |
| − | 5 10 1 0 0 0 | + | 5 10 1 0 0 0 |
| − | 5 11 2 0 0 0 | + | 5 11 2 0 0 0 |
| − | 6 12 2 0 0 0 | + | 6 12 2 0 0 0 |
| − | 7 13 1 1 0 0 | + | 7 13 1 1 0 0 |
| − | 8 14 1 6 0 0 | + | 8 14 1 6 0 0 |
| − | 10 15 1 0 0 0 | + | 10 15 1 0 0 0 |
| − | 13 16 1 0 0 0 | + | 13 16 1 0 0 0 |
| − | 15 17 2 0 0 0 | + | 15 17 2 0 0 0 |
| − | 16 18 1 0 0 0 | + | 16 18 1 0 0 0 |
| − | 18 19 1 0 0 0 | + | 18 19 1 0 0 0 |
| − | 18 20 1 0 0 0 | + | 18 20 1 0 0 0 |
| − | 18 21 2 0 0 0 | + | 18 21 2 0 0 0 |
| − | 19 22 1 0 0 0 | + | 19 22 1 0 0 0 |
| − | 22 23 1 0 0 0 | + | 22 23 1 0 0 0 |
| − | 22 24 1 0 0 0 | + | 22 24 1 0 0 0 |
| − | 22 25 2 0 0 0 | + | 22 25 2 0 0 0 |
| − | 26 23 1 6 0 0 | + | 26 23 1 6 0 0 |
| − | 26 27 1 0 0 0 | + | 26 27 1 0 0 0 |
| − | 26 28 1 0 0 0 | + | 26 28 1 0 0 0 |
| − | 27 29 1 0 0 0 | + | 27 29 1 0 0 0 |
| − | 27 30 1 6 0 0 | + | 27 30 1 6 0 0 |
| − | 28 31 1 0 0 0 | + | 28 31 1 0 0 0 |
| − | 29 32 1 0 0 0 | + | 29 32 1 0 0 0 |
| − | 29 33 1 1 0 0 | + | 29 33 1 1 0 0 |
| − | 30 34 1 0 0 0 | + | 30 34 1 0 0 0 |
| − | 31 35 1 1 0 0 | + | 31 35 1 1 0 0 |
| − | 32 36 1 6 0 0 | + | 32 36 1 6 0 0 |
| − | 34 37 1 0 0 0 | + | 34 37 1 0 0 0 |
| − | 34 38 2 0 0 0 | + | 34 38 2 0 0 0 |
| − | 35 39 1 0 0 0 | + | 35 39 1 0 0 0 |
| − | 7 8 1 0 0 0 | + | 7 8 1 0 0 0 |
| − | 12 15 1 0 0 0 | + | 12 15 1 0 0 0 |
| − | 31 32 1 0 0 0 | + | 31 32 1 0 0 0 |
| − | S SKP | + | S SKP 7 |
| − | CAS_RN 528-04-1 | + | CAS_RN 528-04-1 |
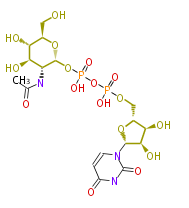
| − | NAME Uridine 5'-diphospho-N-acetylglucosamine | + | NAME Uridine 5'-diphospho-N-acetylglucosamine |
| + | ID PR100620 | ||
| + | FORMULA C17H27N3O17P2 | ||
| + | EXACTMASS 607.081569473 | ||
| + | AVERAGEMASS 607.353822 | ||
| + | SMILES OC[C@@H](O1)[C@@H](O)[C@H](O)[C@@H](NC(C)=O)C1OP(O)(=O)OP(O)(=O)OC[C@@H](O2)[C@@H](O)[C@@H](O)[C@@H]2N(C=3)C(=O)NC(=O)C3 | ||
M END | M END | ||
Latest revision as of 10:28, 23 December 2009
39 41 0 0 1 0 0 0 0 0999 V2000 32.0842 -27.6135 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 33.1497 -25.4069 0.0000 N 0 0 3 0 0 0 0 0 0 0 0 0 30.9722 -26.8043 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 31.6883 -28.8885 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 31.9794 -24.6965 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 34.3492 -24.6965 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 29.8776 -27.5902 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 30.3259 -28.8885 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 32.4918 -29.9948 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 31.9794 -23.3165 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 30.7917 -25.3719 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 34.3492 -23.3165 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 28.5850 -27.1826 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 29.5225 -29.9948 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 33.1556 -22.6411 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 28.2939 -25.8493 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 33.1556 -21.2730 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 26.9256 -25.8493 0.0000 P 0 0 3 0 0 0 0 0 0 0 0 0 25.5632 -25.8493 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 26.9256 -24.4811 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 26.9198 -27.2059 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 24.2066 -25.8493 0.0000 P 0 0 3 0 0 0 0 0 0 0 0 0 22.8325 -25.8435 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 24.2066 -24.4811 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 24.2066 -27.2059 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 21.6506 -26.5130 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 21.6506 -27.8871 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 20.4629 -25.8435 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 20.4629 -28.5625 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 23.0887 -29.0283 0.0000 N 0 0 0 0 0 0 0 0 0 0 0 0 19.2810 -26.5130 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 19.2810 -27.8871 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 20.4629 -29.9249 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 23.5022 -30.2510 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 18.1222 -25.8435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 18.1222 -28.5625 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 24.8471 -30.4198 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 22.6637 -31.3282 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 17.0801 -26.7168 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 1 2 1 1 0 0 1 3 1 0 0 0 1 4 1 0 0 0 2 5 1 0 0 0 2 6 1 0 0 0 3 7 1 0 0 0 4 8 1 0 0 0 4 9 1 6 0 0 5 10 1 0 0 0 5 11 2 0 0 0 6 12 2 0 0 0 7 13 1 1 0 0 8 14 1 6 0 0 10 15 1 0 0 0 13 16 1 0 0 0 15 17 2 0 0 0 16 18 1 0 0 0 18 19 1 0 0 0 18 20 1 0 0 0 18 21 2 0 0 0 19 22 1 0 0 0 22 23 1 0 0 0 22 24 1 0 0 0 22 25 2 0 0 0 26 23 1 6 0 0 26 27 1 0 0 0 26 28 1 0 0 0 27 29 1 0 0 0 27 30 1 6 0 0 28 31 1 0 0 0 29 32 1 0 0 0 29 33 1 1 0 0 30 34 1 0 0 0 31 35 1 1 0 0 32 36 1 6 0 0 34 37 1 0 0 0 34 38 2 0 0 0 35 39 1 0 0 0 7 8 1 0 0 0 12 15 1 0 0 0 31 32 1 0 0 0 S SKP 7 CAS_RN 528-04-1 NAME Uridine 5'-diphospho-N-acetylglucosamine ID PR100620 FORMULA C17H27N3O17P2 EXACTMASS 607.081569473 AVERAGEMASS 607.353822 SMILES OC[C@@H](O1)[C@@H](O)[C@H](O)[C@@H](NC(C)=O)C1OP(O)(=O)OP(O)(=O)OC[C@@H](O2)[C@@H](O)[C@@H](O)[C@@H]2N(C=3)C(=O)NC(=O)C3 M END