Mol:FL5FFAGL0003
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 55 60 0 0 0 0 0 0 0 0999 V2000 | + | 55 60 0 0 0 0 0 0 0 0999 V2000 |
| − | -2.1798 0.5404 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1798 0.5404 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1798 -0.1020 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1798 -0.1020 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6235 -0.4232 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6235 -0.4232 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0671 -0.1020 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0671 -0.1020 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0671 0.5404 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0671 0.5404 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6235 0.8615 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6235 0.8615 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5108 -0.4232 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5108 -0.4232 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0455 -0.1020 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0455 -0.1020 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0455 0.5404 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0455 0.5404 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5108 0.8615 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5108 0.8615 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5108 -0.9240 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5108 -0.9240 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6016 0.8614 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6016 0.8614 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1685 0.5341 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1685 0.5341 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.7355 0.8614 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.7355 0.8614 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.7355 1.5161 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.7355 1.5161 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1685 1.8435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1685 1.8435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6016 1.5161 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6016 1.5161 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.7358 0.8614 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.7358 0.8614 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.4396 -0.4909 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.4396 -0.4909 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6235 -1.0653 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6235 -1.0653 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.3694 1.8821 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.3694 1.8821 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8302 -1.6086 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 0.8302 -1.6086 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 0.2474 -1.2722 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 0.2474 -1.2722 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 0.4323 -1.9192 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.4323 -1.9192 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.2474 -2.5702 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 0.2474 -2.5702 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 0.8302 -2.9068 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 0.8302 -2.9068 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 0.6453 -2.2596 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 0.6453 -2.2596 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 0.6628 -0.9837 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6628 -0.9837 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1068 -2.5261 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1068 -2.5261 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1660 -3.8487 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1660 -3.8487 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8645 -0.3065 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 | + | 1.8645 -0.3065 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 |
| − | 2.3069 -0.8905 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.3069 -0.8905 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 2.9440 -0.6427 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.9440 -0.6427 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 3.5587 -0.6361 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 3.5587 -0.6361 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 3.1120 -0.1893 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 3.1120 -0.1893 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 2.5547 -0.4835 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5547 -0.4835 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.6173 -0.9240 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.6173 -0.9240 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.3090 -1.2555 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.3090 -1.2555 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.5499 -0.7682 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.5499 -0.7682 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.5723 2.7254 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 | + | -3.5723 2.7254 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 |
| − | -3.4888 2.0592 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | -3.4888 2.0592 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | -2.8697 1.9639 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | -2.8697 1.9639 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | -2.3460 1.6538 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | -2.3460 1.6538 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | -2.5285 2.2468 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.5285 2.2468 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.1521 2.3595 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | -3.1521 2.3595 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | -3.8339 3.1787 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.8339 3.1787 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.0722 1.8142 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.0722 1.8142 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.8607 1.3103 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.8607 1.3103 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.6235 1.5037 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.6235 1.5037 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.9520 2.7147 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.9520 2.7147 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.0952 3.5271 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.0952 3.5271 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.1038 -2.7246 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.1038 -2.7246 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.2131 -3.7186 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.2131 -3.7186 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.8689 0.4642 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.8689 0.4642 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.8133 0.1355 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.8133 0.1355 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 1 18 1 0 0 0 0 | + | 1 18 1 0 0 0 0 |
| − | 8 19 1 0 0 0 0 | + | 8 19 1 0 0 0 0 |
| − | 3 20 1 0 0 0 0 | + | 3 20 1 0 0 0 0 |
| − | 21 15 1 0 0 0 0 | + | 21 15 1 0 0 0 0 |
| − | 22 23 1 0 0 0 0 | + | 22 23 1 0 0 0 0 |
| − | 23 24 1 1 0 0 0 | + | 23 24 1 1 0 0 0 |
| − | 24 25 1 1 0 0 0 | + | 24 25 1 1 0 0 0 |
| − | 26 25 1 1 0 0 0 | + | 26 25 1 1 0 0 0 |
| − | 26 27 1 0 0 0 0 | + | 26 27 1 0 0 0 0 |
| − | 27 22 1 0 0 0 0 | + | 27 22 1 0 0 0 0 |
| − | 22 28 1 0 0 0 0 | + | 22 28 1 0 0 0 0 |
| − | 27 29 1 0 0 0 0 | + | 27 29 1 0 0 0 0 |
| − | 26 30 1 0 0 0 0 | + | 26 30 1 0 0 0 0 |
| − | 23 19 1 0 0 0 0 | + | 23 19 1 0 0 0 0 |
| − | 31 32 1 1 0 0 0 | + | 31 32 1 1 0 0 0 |
| − | 32 33 1 1 0 0 0 | + | 32 33 1 1 0 0 0 |
| − | 34 33 1 1 0 0 0 | + | 34 33 1 1 0 0 0 |
| − | 34 35 1 0 0 0 0 | + | 34 35 1 0 0 0 0 |
| − | 35 36 1 0 0 0 0 | + | 35 36 1 0 0 0 0 |
| − | 36 31 1 0 0 0 0 | + | 36 31 1 0 0 0 0 |
| − | 32 37 1 0 0 0 0 | + | 32 37 1 0 0 0 0 |
| − | 33 38 1 0 0 0 0 | + | 33 38 1 0 0 0 0 |
| − | 28 31 1 0 0 0 0 | + | 28 31 1 0 0 0 0 |
| − | 34 39 1 0 0 0 0 | + | 34 39 1 0 0 0 0 |
| − | 40 41 1 1 0 0 0 | + | 40 41 1 1 0 0 0 |
| − | 41 42 1 1 0 0 0 | + | 41 42 1 1 0 0 0 |
| − | 43 42 1 1 0 0 0 | + | 43 42 1 1 0 0 0 |
| − | 43 44 1 0 0 0 0 | + | 43 44 1 0 0 0 0 |
| − | 44 45 1 0 0 0 0 | + | 44 45 1 0 0 0 0 |
| − | 45 40 1 0 0 0 0 | + | 45 40 1 0 0 0 0 |
| − | 40 46 1 0 0 0 0 | + | 40 46 1 0 0 0 0 |
| − | 41 47 1 0 0 0 0 | + | 41 47 1 0 0 0 0 |
| − | 42 48 1 0 0 0 0 | + | 42 48 1 0 0 0 0 |
| − | 6 49 1 0 0 0 0 | + | 6 49 1 0 0 0 0 |
| − | 49 43 1 0 0 0 0 | + | 49 43 1 0 0 0 0 |
| − | 45 50 1 0 0 0 0 | + | 45 50 1 0 0 0 0 |
| − | 50 51 1 0 0 0 0 | + | 50 51 1 0 0 0 0 |
| − | 25 52 1 0 0 0 0 | + | 25 52 1 0 0 0 0 |
| − | 52 53 1 0 0 0 0 | + | 52 53 1 0 0 0 0 |
| − | 35 54 1 0 0 0 0 | + | 35 54 1 0 0 0 0 |
| − | 54 55 1 0 0 0 0 | + | 54 55 1 0 0 0 0 |
| − | M STY 1 3 SUP | + | M STY 1 3 SUP |
| − | M SLB 1 3 3 | + | M SLB 1 3 3 |
| − | M SAL 3 2 54 55 | + | M SAL 3 2 54 55 |
| − | M SBL 3 1 59 | + | M SBL 3 1 59 |
| − | M SMT 3 CH2OH | + | M SMT 3 CH2OH |
| − | M SVB 3 59 3.9322 -0.5128 | + | M SVB 3 59 3.9322 -0.5128 |
| − | M STY 1 2 SUP | + | M STY 1 2 SUP |
| − | M SLB 1 2 2 | + | M SLB 1 2 2 |
| − | M SAL 2 2 52 53 | + | M SAL 2 2 52 53 |
| − | M SBL 2 1 57 | + | M SBL 2 1 57 |
| − | M SMT 2 CH2OH | + | M SMT 2 CH2OH |
| − | M SVB 2 57 -0.0718 -2.7833 | + | M SVB 2 57 -0.0718 -2.7833 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 50 51 | + | M SAL 1 2 50 51 |
| − | M SBL 1 1 55 | + | M SBL 1 1 55 |
| − | M SMT 1 CH2OH | + | M SMT 1 CH2OH |
| − | M SVB 1 55 -3.1919 2.7652 | + | M SVB 1 55 -3.1919 2.7652 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL5FFAGL0003 | + | ID FL5FFAGL0003 |
| − | KNApSAcK_ID C00005353 | + | KNApSAcK_ID C00005353 |
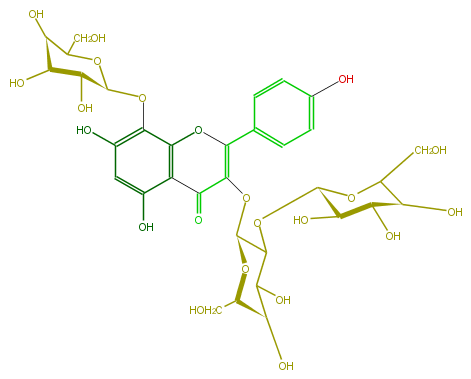
| − | NAME Herbacetin 3-sophoroside-8-glucoside | + | NAME Herbacetin 3-sophoroside-8-glucoside |
| − | CAS_RN 77298-68-1 | + | CAS_RN 77298-68-1 |
| − | FORMULA C33H40O22 | + | FORMULA C33H40O22 |
| − | EXACTMASS 788.201122964 | + | EXACTMASS 788.201122964 |
| − | AVERAGEMASS 788.6575 | + | AVERAGEMASS 788.6575 |
| − | SMILES C(O)C([C@H](O)1)O[C@@H](Oc(c6O)c(c(c(O)c6)3)OC(=C(O[C@H](O5)C(C(O)[C@@H](O)[C@@H](CO)5)O[C@@H](O4)[C@@H]([C@H](O)[C@H](O)C(CO)4)O)C3=O)c(c2)ccc(O)c2)[C@@H](O)[C@H]1O | + | SMILES C(O)C([C@H](O)1)O[C@@H](Oc(c6O)c(c(c(O)c6)3)OC(=C(O[C@H](O5)C(C(O)[C@@H](O)[C@@H](CO)5)O[C@@H](O4)[C@@H]([C@H](O)[C@H](O)C(CO)4)O)C3=O)c(c2)ccc(O)c2)[C@@H](O)[C@H]1O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
55 60 0 0 0 0 0 0 0 0999 V2000
-2.1798 0.5404 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.1798 -0.1020 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6235 -0.4232 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.0671 -0.1020 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.0671 0.5404 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.6235 0.8615 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5108 -0.4232 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0455 -0.1020 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0455 0.5404 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5108 0.8615 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.5108 -0.9240 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.6016 0.8614 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1685 0.5341 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.7355 0.8614 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.7355 1.5161 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1685 1.8435 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.6016 1.5161 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.7358 0.8614 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.4396 -0.4909 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.6235 -1.0653 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.3694 1.8821 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.8302 -1.6086 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
0.2474 -1.2722 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
0.4323 -1.9192 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.2474 -2.5702 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
0.8302 -2.9068 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
0.6453 -2.2596 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
0.6628 -0.9837 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.1068 -2.5261 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.1660 -3.8487 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.8645 -0.3065 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0
2.3069 -0.8905 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
2.9440 -0.6427 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
3.5587 -0.6361 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
3.1120 -0.1893 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
2.5547 -0.4835 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.6173 -0.9240 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.3090 -1.2555 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.5499 -0.7682 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.5723 2.7254 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0
-3.4888 2.0592 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
-2.8697 1.9639 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
-2.3460 1.6538 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
-2.5285 2.2468 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.1521 2.3595 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
-3.8339 3.1787 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.0722 1.8142 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.8607 1.3103 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.6235 1.5037 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.9520 2.7147 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.0952 3.5271 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.1038 -2.7246 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.2131 -3.7186 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.8689 0.4642 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.8133 0.1355 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
1 18 1 0 0 0 0
8 19 1 0 0 0 0
3 20 1 0 0 0 0
21 15 1 0 0 0 0
22 23 1 0 0 0 0
23 24 1 1 0 0 0
24 25 1 1 0 0 0
26 25 1 1 0 0 0
26 27 1 0 0 0 0
27 22 1 0 0 0 0
22 28 1 0 0 0 0
27 29 1 0 0 0 0
26 30 1 0 0 0 0
23 19 1 0 0 0 0
31 32 1 1 0 0 0
32 33 1 1 0 0 0
34 33 1 1 0 0 0
34 35 1 0 0 0 0
35 36 1 0 0 0 0
36 31 1 0 0 0 0
32 37 1 0 0 0 0
33 38 1 0 0 0 0
28 31 1 0 0 0 0
34 39 1 0 0 0 0
40 41 1 1 0 0 0
41 42 1 1 0 0 0
43 42 1 1 0 0 0
43 44 1 0 0 0 0
44 45 1 0 0 0 0
45 40 1 0 0 0 0
40 46 1 0 0 0 0
41 47 1 0 0 0 0
42 48 1 0 0 0 0
6 49 1 0 0 0 0
49 43 1 0 0 0 0
45 50 1 0 0 0 0
50 51 1 0 0 0 0
25 52 1 0 0 0 0
52 53 1 0 0 0 0
35 54 1 0 0 0 0
54 55 1 0 0 0 0
M STY 1 3 SUP
M SLB 1 3 3
M SAL 3 2 54 55
M SBL 3 1 59
M SMT 3 CH2OH
M SVB 3 59 3.9322 -0.5128
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 52 53
M SBL 2 1 57
M SMT 2 CH2OH
M SVB 2 57 -0.0718 -2.7833
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 50 51
M SBL 1 1 55
M SMT 1 CH2OH
M SVB 1 55 -3.1919 2.7652
S SKP 8
ID FL5FFAGL0003
KNApSAcK_ID C00005353
NAME Herbacetin 3-sophoroside-8-glucoside
CAS_RN 77298-68-1
FORMULA C33H40O22
EXACTMASS 788.201122964
AVERAGEMASS 788.6575
SMILES C(O)C([C@H](O)1)O[C@@H](Oc(c6O)c(c(c(O)c6)3)OC(=C(O[C@H](O5)C(C(O)[C@@H](O)[C@@H](CO)5)O[C@@H](O4)[C@@H]([C@H](O)[C@H](O)C(CO)4)O)C3=O)c(c2)ccc(O)c2)[C@@H](O)[C@H]1O
M END