Mol:FL5FECGL0012
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 47 51 0 0 0 0 0 0 0 0999 V2000 | + | 47 51 0 0 0 0 0 0 0 0999 V2000 |
| − | 0.0844 -4.5718 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0844 -4.5718 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7169 -4.4602 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7169 -4.4602 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.9366 -3.8566 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.9366 -3.8566 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5237 -3.3645 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5237 -3.3645 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.1088 -3.4761 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.1088 -3.4761 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.3286 -4.0797 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.3286 -4.0797 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7434 -2.7609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7434 -2.7609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.3305 -2.2688 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.3305 -2.2688 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.3020 -2.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.3020 -2.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5218 -2.9840 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5218 -2.9840 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.2367 -2.6739 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.2367 -2.6739 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.7148 -1.8885 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7148 -1.8885 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.4909 -1.2732 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.4909 -1.2732 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.9117 -0.7717 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.9117 -0.7717 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.5565 -0.8854 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.5565 -0.8854 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.7804 -1.5006 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.7804 -1.5006 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.3596 -2.0022 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.3596 -2.0022 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.1353 -5.1752 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.1353 -5.1752 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5689 -3.7450 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5689 -3.7450 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1993 -0.1194 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1993 -0.1194 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3352 -0.2803 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 1.3352 -0.2803 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 1.0220 -0.8228 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 1.0220 -0.8228 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 1.6243 -0.6507 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.6243 -0.6507 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2303 -0.8228 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.2303 -0.8228 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 2.5436 -0.2803 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.5436 -0.2803 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 1.9412 -0.4524 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 1.9412 -0.4524 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 0.7933 -0.2803 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7933 -0.2803 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.9783 -0.0288 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.9783 -0.0288 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.2216 -0.4620 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.2216 -0.4620 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.7508 -0.5801 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.7508 -0.5801 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.1231 0.0707 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 | + | 5.1231 0.0707 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 |
| − | 4.7468 0.6694 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 4.7468 0.6694 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 4.1135 0.4848 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 4.1135 0.4848 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 3.5223 0.5302 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 3.5223 0.5302 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 3.9139 0.0630 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.9139 0.0630 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.5592 0.2329 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 4.5592 0.2329 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 5.7857 -0.0461 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.7857 -0.0461 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.1946 1.1629 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.1946 1.1629 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.8145 1.1046 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.8145 1.1046 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0665 -0.1207 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0665 -0.1207 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8092 -1.5506 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8092 -1.5506 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6255 -5.1746 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6255 -5.1746 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1559 -5.8065 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1559 -5.8065 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.5346 -0.5764 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.5346 -0.5764 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.9296 -1.4952 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.9296 -1.4952 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.3969 -1.3216 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.3969 -1.3216 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.3573 -1.0427 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.3573 -1.0427 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 1 18 1 0 0 0 0 | + | 1 18 1 0 0 0 0 |
| − | 3 19 1 0 0 0 0 | + | 3 19 1 0 0 0 0 |
| − | 20 15 1 0 0 0 0 | + | 20 15 1 0 0 0 0 |
| − | 21 22 1 0 0 0 0 | + | 21 22 1 0 0 0 0 |
| − | 22 23 1 1 0 0 0 | + | 22 23 1 1 0 0 0 |
| − | 23 24 1 1 0 0 0 | + | 23 24 1 1 0 0 0 |
| − | 25 24 1 1 0 0 0 | + | 25 24 1 1 0 0 0 |
| − | 25 26 1 0 0 0 0 | + | 25 26 1 0 0 0 0 |
| − | 26 21 1 0 0 0 0 | + | 26 21 1 0 0 0 0 |
| − | 21 27 1 0 0 0 0 | + | 21 27 1 0 0 0 0 |
| − | 26 28 1 0 0 0 0 | + | 26 28 1 0 0 0 0 |
| − | 25 29 1 0 0 0 0 | + | 25 29 1 0 0 0 0 |
| − | 24 30 1 0 0 0 0 | + | 24 30 1 0 0 0 0 |
| − | 31 32 1 1 0 0 0 | + | 31 32 1 1 0 0 0 |
| − | 32 33 1 1 0 0 0 | + | 32 33 1 1 0 0 0 |
| − | 34 33 1 1 0 0 0 | + | 34 33 1 1 0 0 0 |
| − | 34 35 1 0 0 0 0 | + | 34 35 1 0 0 0 0 |
| − | 35 36 1 0 0 0 0 | + | 35 36 1 0 0 0 0 |
| − | 36 31 1 0 0 0 0 | + | 36 31 1 0 0 0 0 |
| − | 31 37 1 0 0 0 0 | + | 31 37 1 0 0 0 0 |
| − | 32 38 1 0 0 0 0 | + | 32 38 1 0 0 0 0 |
| − | 33 39 1 0 0 0 0 | + | 33 39 1 0 0 0 0 |
| − | 34 40 1 0 0 0 0 | + | 34 40 1 0 0 0 0 |
| − | 40 30 1 0 0 0 0 | + | 40 30 1 0 0 0 0 |
| − | 22 41 1 0 0 0 0 | + | 22 41 1 0 0 0 0 |
| − | 41 8 1 0 0 0 0 | + | 41 8 1 0 0 0 0 |
| − | 2 42 1 0 0 0 0 | + | 2 42 1 0 0 0 0 |
| − | 42 43 1 0 0 0 0 | + | 42 43 1 0 0 0 0 |
| − | 36 44 1 0 0 0 0 | + | 36 44 1 0 0 0 0 |
| − | 44 45 1 0 0 0 0 | + | 44 45 1 0 0 0 0 |
| − | 16 46 1 0 0 0 0 | + | 16 46 1 0 0 0 0 |
| − | 46 47 1 0 0 0 0 | + | 46 47 1 0 0 0 0 |
| − | M STY 1 3 SUP | + | M STY 1 3 SUP |
| − | M SLB 1 3 3 | + | M SLB 1 3 3 |
| − | M SAL 3 2 44 45 | + | M SAL 3 2 44 45 |
| − | M SBL 3 1 48 | + | M SBL 3 1 48 |
| − | M SMT 3 CH2OH | + | M SMT 3 CH2OH |
| − | M SVB 3 48 1.3577 -1.0393 | + | M SVB 3 48 1.3577 -1.0393 |
| − | M STY 1 2 SUP | + | M STY 1 2 SUP |
| − | M SLB 1 2 2 | + | M SLB 1 2 2 |
| − | M SAL 2 2 46 47 | + | M SAL 2 2 46 47 |
| − | M SBL 2 1 50 | + | M SBL 2 1 50 |
| − | M SMT 2 OCH3 | + | M SMT 2 OCH3 |
| − | M SVB 2 50 1.1245 2.6308 | + | M SVB 2 50 1.1245 2.6308 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 42 43 | + | M SAL 1 2 42 43 |
| − | M SBL 1 1 46 | + | M SBL 1 1 46 |
| − | M SMT 1 OCH3 | + | M SMT 1 OCH3 |
| − | M SVB 1 46 -3.2216 0.2007 | + | M SVB 1 46 -3.2216 0.2007 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL5FECGL0012 | + | ID FL5FECGL0012 |
| − | KNApSAcK_ID C00005668 | + | KNApSAcK_ID C00005668 |
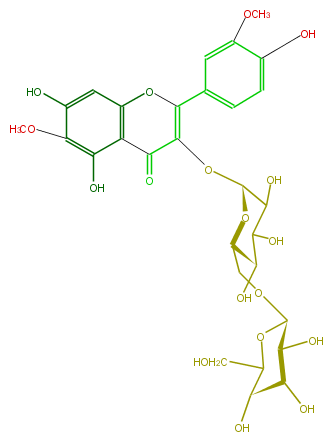
| − | NAME Spinacetin 3-gentiobioside | + | NAME Spinacetin 3-gentiobioside |
| − | CAS_RN 101021-29-8 | + | CAS_RN 101021-29-8 |
| − | FORMULA C29H34O18 | + | FORMULA C29H34O18 |
| − | EXACTMASS 670.174514284 | + | EXACTMASS 670.174514284 |
| − | AVERAGEMASS 670.5694599999999 | + | AVERAGEMASS 670.5694599999999 |
| − | SMILES c(c1)c(O)c(cc(C(=C(O[C@@H](C4O)O[C@H](CO[C@@H]([C@@H](O)5)OC(CO)[C@H](O)[C@@H]5O)[C@@H](C4O)O)2)Oc(c3)c(c(O)c(OC)c(O)3)C2=O)1)OC | + | SMILES c(c1)c(O)c(cc(C(=C(O[C@@H](C4O)O[C@H](CO[C@@H]([C@@H](O)5)OC(CO)[C@H](O)[C@@H]5O)[C@@H](C4O)O)2)Oc(c3)c(c(O)c(OC)c(O)3)C2=O)1)OC |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
47 51 0 0 0 0 0 0 0 0999 V2000
0.0844 -4.5718 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7169 -4.4602 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.9366 -3.8566 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5237 -3.3645 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.1088 -3.4761 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.3286 -4.0797 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7434 -2.7609 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.3305 -2.2688 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.3020 -2.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5218 -2.9840 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.2367 -2.6739 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.7148 -1.8885 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.4909 -1.2732 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.9117 -0.7717 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.5565 -0.8854 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.7804 -1.5006 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.3596 -2.0022 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.1353 -5.1752 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.5689 -3.7450 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.1993 -0.1194 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.3352 -0.2803 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
1.0220 -0.8228 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
1.6243 -0.6507 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.2303 -0.8228 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
2.5436 -0.2803 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
1.9412 -0.4524 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
0.7933 -0.2803 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.9783 -0.0288 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.2216 -0.4620 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.7508 -0.5801 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.1231 0.0707 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0
4.7468 0.6694 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
4.1135 0.4848 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
3.5223 0.5302 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
3.9139 0.0630 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.5592 0.2329 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
5.7857 -0.0461 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.1946 1.1629 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.8145 1.1046 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.0665 -0.1207 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.8092 -1.5506 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.6255 -5.1746 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.1559 -5.8065 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.5346 -0.5764 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.9296 -1.4952 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.3969 -1.3216 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.3573 -1.0427 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
1 18 1 0 0 0 0
3 19 1 0 0 0 0
20 15 1 0 0 0 0
21 22 1 0 0 0 0
22 23 1 1 0 0 0
23 24 1 1 0 0 0
25 24 1 1 0 0 0
25 26 1 0 0 0 0
26 21 1 0 0 0 0
21 27 1 0 0 0 0
26 28 1 0 0 0 0
25 29 1 0 0 0 0
24 30 1 0 0 0 0
31 32 1 1 0 0 0
32 33 1 1 0 0 0
34 33 1 1 0 0 0
34 35 1 0 0 0 0
35 36 1 0 0 0 0
36 31 1 0 0 0 0
31 37 1 0 0 0 0
32 38 1 0 0 0 0
33 39 1 0 0 0 0
34 40 1 0 0 0 0
40 30 1 0 0 0 0
22 41 1 0 0 0 0
41 8 1 0 0 0 0
2 42 1 0 0 0 0
42 43 1 0 0 0 0
36 44 1 0 0 0 0
44 45 1 0 0 0 0
16 46 1 0 0 0 0
46 47 1 0 0 0 0
M STY 1 3 SUP
M SLB 1 3 3
M SAL 3 2 44 45
M SBL 3 1 48
M SMT 3 CH2OH
M SVB 3 48 1.3577 -1.0393
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 46 47
M SBL 2 1 50
M SMT 2 OCH3
M SVB 2 50 1.1245 2.6308
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 42 43
M SBL 1 1 46
M SMT 1 OCH3
M SVB 1 46 -3.2216 0.2007
S SKP 8
ID FL5FECGL0012
KNApSAcK_ID C00005668
NAME Spinacetin 3-gentiobioside
CAS_RN 101021-29-8
FORMULA C29H34O18
EXACTMASS 670.174514284
AVERAGEMASS 670.5694599999999
SMILES c(c1)c(O)c(cc(C(=C(O[C@@H](C4O)O[C@H](CO[C@@H]([C@@H](O)5)OC(CO)[C@H](O)[C@@H]5O)[C@@H](C4O)O)2)Oc(c3)c(c(O)c(OC)c(O)3)C2=O)1)OC
M END