Mol:FL5FECGA0004
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 36 39 0 0 0 0 0 0 0 0999 V2000 | + | 36 39 0 0 0 0 0 0 0 0999 V2000 |
| − | -0.0932 -5.2193 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.0932 -5.2193 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5394 -5.1078 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5394 -5.1078 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7591 -4.5041 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7591 -4.5041 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.3462 -4.0121 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.3462 -4.0121 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.2864 -4.1236 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.2864 -4.1236 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.5061 -4.7272 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.5061 -4.7272 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5659 -3.4083 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5659 -3.4083 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1530 -2.9163 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1530 -2.9163 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.4796 -3.0278 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.4796 -3.0278 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6993 -3.6314 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6993 -3.6314 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.0590 -3.3213 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.0590 -3.3213 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.8924 -2.5359 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.8924 -2.5359 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6685 -1.9208 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6685 -1.9208 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0893 -1.4192 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0893 -1.4192 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.7340 -1.5329 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.7340 -1.5329 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.9579 -2.1482 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.9579 -2.1482 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.5371 -2.6496 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.5371 -2.6496 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5824 -2.1825 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5824 -2.1825 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.0330 -0.8320 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 1.0330 -0.8320 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 0.6454 -1.5034 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 0.6454 -1.5034 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 1.3908 -1.2904 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3908 -1.2904 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.1408 -1.5034 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.1408 -1.5034 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 2.5285 -0.8320 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.5285 -0.8320 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 1.7830 -1.0450 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 1.7830 -1.0450 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 0.3059 -1.0268 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.3059 -1.0268 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0518 -0.5794 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0518 -0.5794 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.3052 0.0014 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.3052 0.0014 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1547 -1.0316 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1547 -1.0316 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3913 -4.3926 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3913 -4.3926 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.6024 -2.2618 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.6024 -2.2618 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.8858 -1.4143 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.8858 -1.4143 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.6002 -1.8268 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.6002 -1.8268 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.4480 -5.8222 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.4480 -5.8222 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6616 -6.6190 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6616 -6.6190 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6405 -5.6785 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6405 -5.6785 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4066 -6.3213 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4066 -6.3213 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 19 20 1 0 0 0 0 | + | 19 20 1 0 0 0 0 |
| − | 20 21 1 1 0 0 0 | + | 20 21 1 1 0 0 0 |
| − | 23 22 1 1 0 0 0 | + | 23 22 1 1 0 0 0 |
| − | 23 24 1 0 0 0 0 | + | 23 24 1 0 0 0 0 |
| − | 24 19 1 0 0 0 0 | + | 24 19 1 0 0 0 0 |
| − | 19 25 1 0 0 0 0 | + | 19 25 1 0 0 0 0 |
| − | 24 26 1 0 0 0 0 | + | 24 26 1 0 0 0 0 |
| − | 23 27 1 0 0 0 0 | + | 23 27 1 0 0 0 0 |
| − | 20 18 1 0 0 0 0 | + | 20 18 1 0 0 0 0 |
| − | 18 8 1 0 0 0 0 | + | 18 8 1 0 0 0 0 |
| − | 22 21 1 1 0 0 0 | + | 22 21 1 1 0 0 0 |
| − | 15 28 1 0 0 0 0 | + | 15 28 1 0 0 0 0 |
| − | 3 29 1 0 0 0 0 | + | 3 29 1 0 0 0 0 |
| − | 16 30 1 0 0 0 0 | + | 16 30 1 0 0 0 0 |
| − | 22 31 1 0 0 0 0 | + | 22 31 1 0 0 0 0 |
| − | 31 32 1 0 0 0 0 | + | 31 32 1 0 0 0 0 |
| − | 2 33 1 0 0 0 0 | + | 2 33 1 0 0 0 0 |
| − | 33 34 1 0 0 0 0 | + | 33 34 1 0 0 0 0 |
| − | 1 35 1 0 0 0 0 | + | 1 35 1 0 0 0 0 |
| − | 35 36 1 0 0 0 0 | + | 35 36 1 0 0 0 0 |
| − | M STY 1 3 SUP | + | M STY 1 3 SUP |
| − | M SLB 1 3 3 | + | M SLB 1 3 3 |
| − | M SAL 3 2 31 32 | + | M SAL 3 2 31 32 |
| − | M SBL 3 1 34 | + | M SBL 3 1 34 |
| − | M SMT 3 CH2OH | + | M SMT 3 CH2OH |
| − | M SVB 3 34 2.8858 -1.4143 | + | M SVB 3 34 2.8858 -1.4143 |
| − | M STY 1 2 SUP | + | M STY 1 2 SUP |
| − | M SLB 1 2 2 | + | M SLB 1 2 2 |
| − | M SAL 2 2 35 36 | + | M SAL 2 2 35 36 |
| − | M SBL 2 1 38 | + | M SBL 2 1 38 |
| − | M SMT 2 OCH3 | + | M SMT 2 OCH3 |
| − | M SVB 2 38 -3.243 0.488 | + | M SVB 2 38 -3.243 0.488 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 33 34 | + | M SAL 1 2 33 34 |
| − | M SBL 1 1 36 | + | M SBL 1 1 36 |
| − | M SMT 1 OCH3 | + | M SMT 1 OCH3 |
| − | M SVB 1 36 -3.6002 -0.6817 | + | M SVB 1 36 -3.6002 -0.6817 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL5FECGA0004 | + | ID FL5FECGA0004 |
| − | KNApSAcK_ID C00005663 | + | KNApSAcK_ID C00005663 |
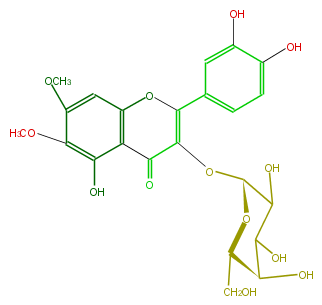
| − | NAME Eupatolitin 3-galactoside | + | NAME Eupatolitin 3-galactoside |
| − | CAS_RN 98604-36-5 | + | CAS_RN 98604-36-5 |
| − | FORMULA C23H24O13 | + | FORMULA C23H24O13 |
| − | EXACTMASS 508.121690854 | + | EXACTMASS 508.121690854 |
| − | AVERAGEMASS 508.42886 | + | AVERAGEMASS 508.42886 |
| − | SMILES O(C(=C(c(c4)cc(c(O)c4)O)3)C(=O)c(c2O)c(O3)cc(c2OC)OC)[C@H](O1)C(O)C([C@H]([C@@H](CO)1)O)O | + | SMILES O(C(=C(c(c4)cc(c(O)c4)O)3)C(=O)c(c2O)c(O3)cc(c2OC)OC)[C@H](O1)C(O)C([C@H]([C@@H](CO)1)O)O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
36 39 0 0 0 0 0 0 0 0999 V2000
-0.0932 -5.2193 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5394 -5.1078 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7591 -4.5041 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.3462 -4.0121 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.2864 -4.1236 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.5061 -4.7272 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5659 -3.4083 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.1530 -2.9163 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.4796 -3.0278 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.6993 -3.6314 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.0590 -3.3213 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.8924 -2.5359 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.6685 -1.9208 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.0893 -1.4192 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.7340 -1.5329 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.9579 -2.1482 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.5371 -2.6496 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.5824 -2.1825 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.0330 -0.8320 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
0.6454 -1.5034 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
1.3908 -1.2904 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.1408 -1.5034 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
2.5285 -0.8320 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
1.7830 -1.0450 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
0.3059 -1.0268 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.0518 -0.5794 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.3052 0.0014 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.1547 -1.0316 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.3913 -4.3926 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.6024 -2.2618 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.8858 -1.4143 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.6002 -1.8268 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.4480 -5.8222 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.6616 -6.6190 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.6405 -5.6785 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-1.4066 -6.3213 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
19 20 1 0 0 0 0
20 21 1 1 0 0 0
23 22 1 1 0 0 0
23 24 1 0 0 0 0
24 19 1 0 0 0 0
19 25 1 0 0 0 0
24 26 1 0 0 0 0
23 27 1 0 0 0 0
20 18 1 0 0 0 0
18 8 1 0 0 0 0
22 21 1 1 0 0 0
15 28 1 0 0 0 0
3 29 1 0 0 0 0
16 30 1 0 0 0 0
22 31 1 0 0 0 0
31 32 1 0 0 0 0
2 33 1 0 0 0 0
33 34 1 0 0 0 0
1 35 1 0 0 0 0
35 36 1 0 0 0 0
M STY 1 3 SUP
M SLB 1 3 3
M SAL 3 2 31 32
M SBL 3 1 34
M SMT 3 CH2OH
M SVB 3 34 2.8858 -1.4143
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 35 36
M SBL 2 1 38
M SMT 2 OCH3
M SVB 2 38 -3.243 0.488
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 33 34
M SBL 1 1 36
M SMT 1 OCH3
M SVB 1 36 -3.6002 -0.6817
S SKP 8
ID FL5FECGA0004
KNApSAcK_ID C00005663
NAME Eupatolitin 3-galactoside
CAS_RN 98604-36-5
FORMULA C23H24O13
EXACTMASS 508.121690854
AVERAGEMASS 508.42886
SMILES O(C(=C(c(c4)cc(c(O)c4)O)3)C(=O)c(c2O)c(O3)cc(c2OC)OC)[C@H](O1)C(O)C([C@H]([C@@H](CO)1)O)O
M END