Mol:FL5FADGA0015
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 56 61 0 0 0 0 0 0 0 0999 V2000 | + | 56 61 0 0 0 0 0 0 0 0999 V2000 |
| − | -1.4723 0.9817 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4723 0.9817 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4723 0.3393 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4723 0.3393 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.9160 0.0181 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.9160 0.0181 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.3597 0.3393 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.3597 0.3393 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.3597 0.9817 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.3597 0.9817 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.9160 1.3028 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.9160 1.3028 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1966 0.0181 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1966 0.0181 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7529 0.3393 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7529 0.3393 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7529 0.9817 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7529 0.9817 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1966 1.3028 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1966 1.3028 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1966 -0.4827 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1966 -0.4827 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3090 1.3027 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3090 1.3027 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8759 0.9754 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8759 0.9754 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.4429 1.3027 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.4429 1.3027 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.4429 1.9574 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.4429 1.9574 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8759 2.2847 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8759 2.2847 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3090 1.9574 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3090 1.9574 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.0284 1.3027 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.0284 1.3027 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1470 -0.0496 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1470 -0.0496 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.9160 -0.6240 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.9160 -0.6240 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.3089 2.4574 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.3089 2.4574 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5376 -1.1674 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 1.5376 -1.1674 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 0.9548 -0.8309 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 0.9548 -0.8309 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 1.1398 -1.4779 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1398 -1.4779 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.9548 -2.1289 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 0.9548 -2.1289 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 1.5376 -2.4655 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 1.5376 -2.4655 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 1.3527 -1.8184 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 1.3527 -1.8184 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 1.3702 -0.5424 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3702 -0.5424 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.7840 -1.9339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.7840 -1.9339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.3210 -2.0132 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.3210 -2.0132 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.3038 0.2173 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 | + | 2.3038 0.2173 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 |
| − | 2.7462 -0.3667 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 2.7462 -0.3667 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 3.3833 -0.1189 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 3.3833 -0.1189 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 3.9980 -0.1123 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | 3.9980 -0.1123 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | 3.5513 0.3345 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | 3.5513 0.3345 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | 2.9940 0.0403 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.9940 0.0403 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0565 -0.4002 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0565 -0.4002 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.7483 -0.7317 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.7483 -0.7317 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.6845 -0.3622 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.6845 -0.3622 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.2264 1.1344 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 | + | -4.2264 1.1344 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0 |
| − | -3.7994 0.5708 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | -3.7994 0.5708 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | -3.1847 0.8099 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | -3.1847 0.8099 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | -2.5914 0.8163 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 | + | -2.5914 0.8163 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0 |
| − | -3.0225 1.2475 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.0225 1.2475 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.6505 1.0220 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 | + | -3.6505 1.0220 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0 |
| − | -4.7753 1.1824 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.7753 1.1824 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.2024 0.0402 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.2024 0.0402 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.8324 0.2186 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.8324 0.2186 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.8857 1.7478 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.8857 1.7478 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.8637 1.9565 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.8637 1.9565 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.5980 -2.1498 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.5980 -2.1498 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.1163 -2.5621 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.1163 -2.5621 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.1593 2.8609 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.1593 2.8609 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.6006 3.7582 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.6006 3.7582 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.3173 0.7544 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.3173 0.7544 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.3395 1.7541 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.3395 1.7541 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 3 4 1 0 0 0 0 | + | 3 4 1 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 1 18 1 0 0 0 0 | + | 1 18 1 0 0 0 0 |
| − | 8 19 1 0 0 0 0 | + | 8 19 1 0 0 0 0 |
| − | 3 20 1 0 0 0 0 | + | 3 20 1 0 0 0 0 |
| − | 21 15 1 0 0 0 0 | + | 21 15 1 0 0 0 0 |
| − | 22 23 1 0 0 0 0 | + | 22 23 1 0 0 0 0 |
| − | 23 24 1 1 0 0 0 | + | 23 24 1 1 0 0 0 |
| − | 24 25 1 1 0 0 0 | + | 24 25 1 1 0 0 0 |
| − | 26 25 1 1 0 0 0 | + | 26 25 1 1 0 0 0 |
| − | 26 27 1 0 0 0 0 | + | 26 27 1 0 0 0 0 |
| − | 27 22 1 0 0 0 0 | + | 27 22 1 0 0 0 0 |
| − | 22 28 1 0 0 0 0 | + | 22 28 1 0 0 0 0 |
| − | 27 29 1 0 0 0 0 | + | 27 29 1 0 0 0 0 |
| − | 26 30 1 0 0 0 0 | + | 26 30 1 0 0 0 0 |
| − | 23 19 1 0 0 0 0 | + | 23 19 1 0 0 0 0 |
| − | 31 32 1 1 0 0 0 | + | 31 32 1 1 0 0 0 |
| − | 32 33 1 1 0 0 0 | + | 32 33 1 1 0 0 0 |
| − | 34 33 1 1 0 0 0 | + | 34 33 1 1 0 0 0 |
| − | 34 35 1 0 0 0 0 | + | 34 35 1 0 0 0 0 |
| − | 35 36 1 0 0 0 0 | + | 35 36 1 0 0 0 0 |
| − | 36 31 1 0 0 0 0 | + | 36 31 1 0 0 0 0 |
| − | 32 37 1 0 0 0 0 | + | 32 37 1 0 0 0 0 |
| − | 33 38 1 0 0 0 0 | + | 33 38 1 0 0 0 0 |
| − | 31 28 1 0 0 0 0 | + | 31 28 1 0 0 0 0 |
| − | 34 39 1 0 0 0 0 | + | 34 39 1 0 0 0 0 |
| − | 40 41 1 1 0 0 0 | + | 40 41 1 1 0 0 0 |
| − | 41 42 1 1 0 0 0 | + | 41 42 1 1 0 0 0 |
| − | 43 42 1 1 0 0 0 | + | 43 42 1 1 0 0 0 |
| − | 43 44 1 0 0 0 0 | + | 43 44 1 0 0 0 0 |
| − | 44 45 1 0 0 0 0 | + | 44 45 1 0 0 0 0 |
| − | 45 40 1 0 0 0 0 | + | 45 40 1 0 0 0 0 |
| − | 40 46 1 0 0 0 0 | + | 40 46 1 0 0 0 0 |
| − | 41 47 1 0 0 0 0 | + | 41 47 1 0 0 0 0 |
| − | 42 48 1 0 0 0 0 | + | 42 48 1 0 0 0 0 |
| − | 43 18 1 0 0 0 0 | + | 43 18 1 0 0 0 0 |
| − | 45 49 1 0 0 0 0 | + | 45 49 1 0 0 0 0 |
| − | 49 50 1 0 0 0 0 | + | 49 50 1 0 0 0 0 |
| − | 25 51 1 0 0 0 0 | + | 25 51 1 0 0 0 0 |
| − | 51 52 1 0 0 0 0 | + | 51 52 1 0 0 0 0 |
| − | 16 53 1 0 0 0 0 | + | 16 53 1 0 0 0 0 |
| − | 53 54 1 0 0 0 0 | + | 53 54 1 0 0 0 0 |
| − | 35 55 1 0 0 0 0 | + | 35 55 1 0 0 0 0 |
| − | 55 56 1 0 0 0 0 | + | 55 56 1 0 0 0 0 |
| − | M STY 1 4 SUP | + | M STY 1 4 SUP |
| − | M SLB 1 4 4 | + | M SLB 1 4 4 |
| − | M SAL 4 2 55 56 | + | M SAL 4 2 55 56 |
| − | M SBL 4 1 60 | + | M SBL 4 1 60 |
| − | M SMT 4 CH2OH | + | M SMT 4 CH2OH |
| − | M SVB 4 60 4.3653 0.0176 | + | M SVB 4 60 4.3653 0.0176 |
| − | M STY 1 3 SUP | + | M STY 1 3 SUP |
| − | M SLB 1 3 3 | + | M SLB 1 3 3 |
| − | M SAL 3 2 51 52 | + | M SAL 3 2 51 52 |
| − | M SBL 3 1 56 | + | M SBL 3 1 56 |
| − | M SMT 3 CH2OH | + | M SMT 3 CH2OH |
| − | M SVB 3 56 0.7944 -2.4484 | + | M SVB 3 56 0.7944 -2.4484 |
| − | M STY 1 2 SUP | + | M STY 1 2 SUP |
| − | M SLB 1 2 2 | + | M SLB 1 2 2 |
| − | M SAL 2 2 49 50 | + | M SAL 2 2 49 50 |
| − | M SBL 2 1 54 | + | M SBL 2 1 54 |
| − | M SMT 2 CH2OH | + | M SMT 2 CH2OH |
| − | M SVB 2 54 -3.8857 1.7478 | + | M SVB 2 54 -3.8857 1.7478 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 53 54 | + | M SAL 1 2 53 54 |
| − | M SBL 1 1 58 | + | M SBL 1 1 58 |
| − | M SMT 1 OCH3 | + | M SMT 1 OCH3 |
| − | M SVB 1 58 2.1593 2.8609 | + | M SVB 1 58 2.1593 2.8609 |
| − | S SKP 8 | + | S SKP 8 |
| − | ID FL5FADGA0015 | + | ID FL5FADGA0015 |
| − | KNApSAcK_ID C00005574 | + | KNApSAcK_ID C00005574 |
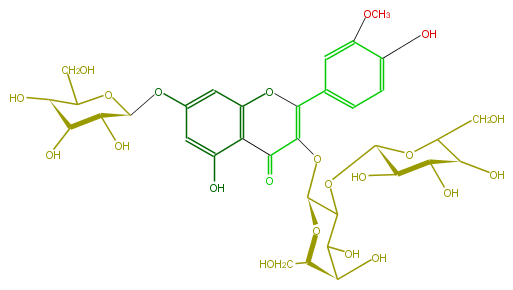
| − | NAME Isorhamnetin 3-glucosyl-(1->2)-galactoside-7-glucoside | + | NAME Isorhamnetin 3-glucosyl-(1->2)-galactoside-7-glucoside |
| − | CAS_RN - | + | CAS_RN - |
| − | FORMULA C34H42O22 | + | FORMULA C34H42O22 |
| − | EXACTMASS 802.216773028 | + | EXACTMASS 802.216773028 |
| − | AVERAGEMASS 802.68408 | + | AVERAGEMASS 802.68408 |
| − | SMILES C(C([C@H]6O)O[C@@H]([C@@H]([C@@H]6O)O)OC([C@H](OC(=C2c(c5)cc(c(c5)O)OC)C(c(c4O)c(cc(c4)O[C@@H]([C@@H](O)3)OC([C@@H]([C@H](O)3)O)CO)O2)=O)1)C([C@H]([C@@H](CO)O1)O)O)O | + | SMILES C(C([C@H]6O)O[C@@H]([C@@H]([C@@H]6O)O)OC([C@H](OC(=C2c(c5)cc(c(c5)O)OC)C(c(c4O)c(cc(c4)O[C@@H]([C@@H](O)3)OC([C@@H]([C@H](O)3)O)CO)O2)=O)1)C([C@H]([C@@H](CO)O1)O)O)O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
56 61 0 0 0 0 0 0 0 0999 V2000
-1.4723 0.9817 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.4723 0.3393 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.9160 0.0181 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.3597 0.3393 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.3597 0.9817 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.9160 1.3028 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.1966 0.0181 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7529 0.3393 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7529 0.9817 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.1966 1.3028 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.1966 -0.4827 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.3090 1.3027 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8759 0.9754 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.4429 1.3027 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.4429 1.9574 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.8759 2.2847 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.3090 1.9574 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.0284 1.3027 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.1470 -0.0496 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.9160 -0.6240 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.3089 2.4574 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.5376 -1.1674 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
0.9548 -0.8309 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
1.1398 -1.4779 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.9548 -2.1289 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
1.5376 -2.4655 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
1.3527 -1.8184 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
1.3702 -0.5424 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.7840 -1.9339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.3210 -2.0132 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.3038 0.2173 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0
2.7462 -0.3667 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
3.3833 -0.1189 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
3.9980 -0.1123 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
3.5513 0.3345 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
2.9940 0.0403 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.0565 -0.4002 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
3.7483 -0.7317 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.6845 -0.3622 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.2264 1.1344 0.0000 C 0 0 2 0 0 0 0 0 0 0 0 0
-3.7994 0.5708 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
-3.1847 0.8099 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
-2.5914 0.8163 0.0000 C 0 0 1 0 0 0 0 0 0 0 0 0
-3.0225 1.2475 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.6505 1.0220 0.0000 C 0 0 3 0 0 0 0 0 0 0 0 0
-4.7753 1.1824 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.2024 0.0402 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.8324 0.2186 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.8857 1.7478 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.8637 1.9565 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.5980 -2.1498 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.1163 -2.5621 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.1593 2.8609 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.6006 3.7582 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3173 0.7544 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3395 1.7541 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
3 4 1 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
1 18 1 0 0 0 0
8 19 1 0 0 0 0
3 20 1 0 0 0 0
21 15 1 0 0 0 0
22 23 1 0 0 0 0
23 24 1 1 0 0 0
24 25 1 1 0 0 0
26 25 1 1 0 0 0
26 27 1 0 0 0 0
27 22 1 0 0 0 0
22 28 1 0 0 0 0
27 29 1 0 0 0 0
26 30 1 0 0 0 0
23 19 1 0 0 0 0
31 32 1 1 0 0 0
32 33 1 1 0 0 0
34 33 1 1 0 0 0
34 35 1 0 0 0 0
35 36 1 0 0 0 0
36 31 1 0 0 0 0
32 37 1 0 0 0 0
33 38 1 0 0 0 0
31 28 1 0 0 0 0
34 39 1 0 0 0 0
40 41 1 1 0 0 0
41 42 1 1 0 0 0
43 42 1 1 0 0 0
43 44 1 0 0 0 0
44 45 1 0 0 0 0
45 40 1 0 0 0 0
40 46 1 0 0 0 0
41 47 1 0 0 0 0
42 48 1 0 0 0 0
43 18 1 0 0 0 0
45 49 1 0 0 0 0
49 50 1 0 0 0 0
25 51 1 0 0 0 0
51 52 1 0 0 0 0
16 53 1 0 0 0 0
53 54 1 0 0 0 0
35 55 1 0 0 0 0
55 56 1 0 0 0 0
M STY 1 4 SUP
M SLB 1 4 4
M SAL 4 2 55 56
M SBL 4 1 60
M SMT 4 CH2OH
M SVB 4 60 4.3653 0.0176
M STY 1 3 SUP
M SLB 1 3 3
M SAL 3 2 51 52
M SBL 3 1 56
M SMT 3 CH2OH
M SVB 3 56 0.7944 -2.4484
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 49 50
M SBL 2 1 54
M SMT 2 CH2OH
M SVB 2 54 -3.8857 1.7478
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 53 54
M SBL 1 1 58
M SMT 1 OCH3
M SVB 1 58 2.1593 2.8609
S SKP 8
ID FL5FADGA0015
KNApSAcK_ID C00005574
NAME Isorhamnetin 3-glucosyl-(1->2)-galactoside-7-glucoside
CAS_RN -
FORMULA C34H42O22
EXACTMASS 802.216773028
AVERAGEMASS 802.68408
SMILES C(C([C@H]6O)O[C@@H]([C@@H]([C@@H]6O)O)OC([C@H](OC(=C2c(c5)cc(c(c5)O)OC)C(c(c4O)c(cc(c4)O[C@@H]([C@@H](O)3)OC([C@@H]([C@H](O)3)O)CO)O2)=O)1)C([C@H]([C@@H](CO)O1)O)O)O
M END