Mol:FL5FACGL0048
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 54 59 0 0 0 0 0 0 0 0999 V2000 | + | 54 59 0 0 0 0 0 0 0 0999 V2000 |
| − | -1.7500 -0.8707 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.7500 -0.8707 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.7500 -1.6956 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.7500 -1.6956 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0356 -2.1082 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0356 -2.1082 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.3211 -1.6956 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.3211 -1.6956 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.3211 -0.8707 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.3211 -0.8707 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0356 -0.4582 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0356 -0.4582 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.3933 -2.1082 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.3933 -2.1082 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1078 -1.6956 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1078 -1.6956 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.1078 -0.8707 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.1078 -0.8707 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.3933 -0.4582 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.3933 -0.4582 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.3933 -2.7515 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.3933 -2.7515 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0074 -0.2995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0074 -0.2995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.7356 -0.7198 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.7356 -0.7198 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.4639 -0.2995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.4639 -0.2995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.4639 0.5414 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.4639 0.5414 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.7356 0.9618 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.7356 0.9618 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.0074 0.5414 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.0074 0.5414 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.0356 -2.9328 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.0356 -2.9328 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.2432 0.9914 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.2432 0.9914 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.8739 -2.2231 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.8739 -2.2231 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.7356 1.8024 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.7356 1.8024 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.9310 -1.7150 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.9310 -1.7150 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.5447 -2.3841 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.5447 -2.3841 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.2875 -2.1718 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.2875 -2.1718 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.0350 -2.3841 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.0350 -2.3841 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.4215 -1.7150 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.4215 -1.7150 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.6785 -1.9273 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.6785 -1.9273 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.1323 -1.7643 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.1323 -1.7643 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.2745 -1.3163 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.2745 -1.3163 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.1041 -1.7907 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.1041 -1.7907 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.1657 -0.2865 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.1657 -0.2865 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.6227 -1.0032 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.6227 -1.0032 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.8406 -0.6992 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.8406 -0.6992 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.0861 -0.6910 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.0861 -0.6910 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.6344 -0.1426 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.6344 -0.1426 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.3185 -0.5038 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.3185 -0.5038 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.7000 -0.1059 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.7000 -0.1059 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.7097 -0.5363 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.7097 -0.5363 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.2821 -1.0252 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.2821 -1.0252 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.2057 -1.1639 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.2057 -1.1639 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.2272 0.3740 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.2272 0.3740 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.5414 2.6247 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.5414 2.6247 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.7946 1.7714 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.7946 1.7714 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.1103 1.4091 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.1103 1.4091 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.6265 0.7307 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.6265 0.7307 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.4729 1.5478 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.4729 1.5478 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.1088 1.8532 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.1088 1.8532 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.1084 2.9328 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.1084 2.9328 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -5.1598 2.2855 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -5.1598 2.2855 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.0552 2.6158 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.0552 2.6158 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.1373 2.1422 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.1373 2.1422 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.4888 -0.4442 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.4888 -0.4442 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.7921 -2.2167 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.7921 -2.2167 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 5.7097 -2.7465 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 5.7097 -2.7465 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 3 18 1 0 0 0 0 | + | 3 18 1 0 0 0 0 |
| − | 19 15 1 0 0 0 0 | + | 19 15 1 0 0 0 0 |
| − | 20 8 1 0 0 0 0 | + | 20 8 1 0 0 0 0 |
| − | 4 3 1 0 0 0 0 | + | 4 3 1 0 0 0 0 |
| − | 16 21 1 0 0 0 0 | + | 16 21 1 0 0 0 0 |
| − | 22 23 1 0 0 0 0 | + | 22 23 1 0 0 0 0 |
| − | 23 24 1 1 0 0 0 | + | 23 24 1 1 0 0 0 |
| − | 24 25 1 1 0 0 0 | + | 24 25 1 1 0 0 0 |
| − | 26 25 1 1 0 0 0 | + | 26 25 1 1 0 0 0 |
| − | 26 27 1 0 0 0 0 | + | 26 27 1 0 0 0 0 |
| − | 27 22 1 0 0 0 0 | + | 27 22 1 0 0 0 0 |
| − | 22 28 1 0 0 0 0 | + | 22 28 1 0 0 0 0 |
| − | 27 29 1 0 0 0 0 | + | 27 29 1 0 0 0 0 |
| − | 26 30 1 0 0 0 0 | + | 26 30 1 0 0 0 0 |
| − | 23 20 1 0 0 0 0 | + | 23 20 1 0 0 0 0 |
| − | 31 32 1 1 0 0 0 | + | 31 32 1 1 0 0 0 |
| − | 32 33 1 1 0 0 0 | + | 32 33 1 1 0 0 0 |
| − | 34 33 1 1 0 0 0 | + | 34 33 1 1 0 0 0 |
| − | 34 35 1 0 0 0 0 | + | 34 35 1 0 0 0 0 |
| − | 35 36 1 0 0 0 0 | + | 35 36 1 0 0 0 0 |
| − | 36 31 1 0 0 0 0 | + | 36 31 1 0 0 0 0 |
| − | 36 37 1 0 0 0 0 | + | 36 37 1 0 0 0 0 |
| − | 31 38 1 0 0 0 0 | + | 31 38 1 0 0 0 0 |
| − | 32 39 1 0 0 0 0 | + | 32 39 1 0 0 0 0 |
| − | 33 40 1 0 0 0 0 | + | 33 40 1 0 0 0 0 |
| − | 41 37 1 0 0 0 0 | + | 41 37 1 0 0 0 0 |
| − | 42 43 1 1 0 0 0 | + | 42 43 1 1 0 0 0 |
| − | 43 44 1 1 0 0 0 | + | 43 44 1 1 0 0 0 |
| − | 45 44 1 1 0 0 0 | + | 45 44 1 1 0 0 0 |
| − | 45 46 1 0 0 0 0 | + | 45 46 1 0 0 0 0 |
| − | 46 47 1 0 0 0 0 | + | 46 47 1 0 0 0 0 |
| − | 47 42 1 0 0 0 0 | + | 47 42 1 0 0 0 0 |
| − | 45 41 1 0 0 0 0 | + | 45 41 1 0 0 0 0 |
| − | 42 48 1 0 0 0 0 | + | 42 48 1 0 0 0 0 |
| − | 43 49 1 0 0 0 0 | + | 43 49 1 0 0 0 0 |
| − | 47 50 1 0 0 0 0 | + | 47 50 1 0 0 0 0 |
| − | 46 51 1 0 0 0 0 | + | 46 51 1 0 0 0 0 |
| − | 1 52 1 0 0 0 0 | + | 1 52 1 0 0 0 0 |
| − | 52 34 1 0 0 0 0 | + | 52 34 1 0 0 0 0 |
| − | 53 54 1 0 0 0 0 | + | 53 54 1 0 0 0 0 |
| − | 25 53 1 0 0 0 0 | + | 25 53 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 53 54 | + | M SAL 1 2 53 54 |
| − | M SBL 1 1 59 | + | M SBL 1 1 59 |
| − | M SMT 1 CH2OH | + | M SMT 1 CH2OH |
| − | M SBV 1 59 -0.7571 -0.1674 | + | M SBV 1 59 -0.7571 -0.1674 |
| − | S SKP 5 | + | S SKP 5 |
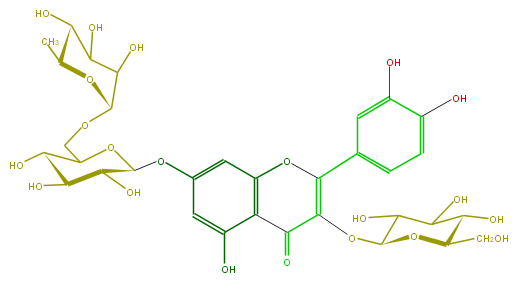
| − | ID FL5FACGL0048 | + | ID FL5FACGL0048 |
| − | FORMULA C33H40O21 | + | FORMULA C33H40O21 |
| − | EXACTMASS 772.206208342 | + | EXACTMASS 772.206208342 |
| − | AVERAGEMASS 772.6581 | + | AVERAGEMASS 772.6581 |
| − | SMILES C(O6)(C(O)C(C(O)C6CO)O)OC(=C(c(c5)ccc(c5O)O)4)C(c(c(O)3)c(O4)cc(c3)OC(C1O)OC(COC(C2O)OC(C)C(C(O)2)O)C(C1O)O)=O | + | SMILES C(O6)(C(O)C(C(O)C6CO)O)OC(=C(c(c5)ccc(c5O)O)4)C(c(c(O)3)c(O4)cc(c3)OC(C1O)OC(COC(C2O)OC(C)C(C(O)2)O)C(C1O)O)=O |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
54 59 0 0 0 0 0 0 0 0999 V2000
-1.7500 -0.8707 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.7500 -1.6956 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.0356 -2.1082 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.3211 -1.6956 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.3211 -0.8707 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.0356 -0.4582 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.3933 -2.1082 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1078 -1.6956 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.1078 -0.8707 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.3933 -0.4582 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.3933 -2.7515 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.0074 -0.2995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.7356 -0.7198 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.4639 -0.2995 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.4639 0.5414 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.7356 0.9618 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.0074 0.5414 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.0356 -2.9328 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.2432 0.9914 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.8739 -2.2231 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.7356 1.8024 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.9310 -1.7150 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.5447 -2.3841 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.2875 -2.1718 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.0350 -2.3841 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.4215 -1.7150 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.6785 -1.9273 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.1323 -1.7643 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.2745 -1.3163 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
5.1041 -1.7907 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-5.1657 -0.2865 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.6227 -1.0032 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.8406 -0.6992 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.0861 -0.6910 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.6344 -0.1426 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.3185 -0.5038 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.7000 -0.1059 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.7097 -0.5363 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-5.2821 -1.0252 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.2057 -1.1639 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.2272 0.3740 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.5414 2.6247 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.7946 1.7714 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.1103 1.4091 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.6265 0.7307 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.4729 1.5478 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.1088 1.8532 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-5.1084 2.9328 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-5.1598 2.2855 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.0552 2.6158 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-3.1373 2.1422 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.4888 -0.4442 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.7921 -2.2167 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
5.7097 -2.7465 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
3 18 1 0 0 0 0
19 15 1 0 0 0 0
20 8 1 0 0 0 0
4 3 1 0 0 0 0
16 21 1 0 0 0 0
22 23 1 0 0 0 0
23 24 1 1 0 0 0
24 25 1 1 0 0 0
26 25 1 1 0 0 0
26 27 1 0 0 0 0
27 22 1 0 0 0 0
22 28 1 0 0 0 0
27 29 1 0 0 0 0
26 30 1 0 0 0 0
23 20 1 0 0 0 0
31 32 1 1 0 0 0
32 33 1 1 0 0 0
34 33 1 1 0 0 0
34 35 1 0 0 0 0
35 36 1 0 0 0 0
36 31 1 0 0 0 0
36 37 1 0 0 0 0
31 38 1 0 0 0 0
32 39 1 0 0 0 0
33 40 1 0 0 0 0
41 37 1 0 0 0 0
42 43 1 1 0 0 0
43 44 1 1 0 0 0
45 44 1 1 0 0 0
45 46 1 0 0 0 0
46 47 1 0 0 0 0
47 42 1 0 0 0 0
45 41 1 0 0 0 0
42 48 1 0 0 0 0
43 49 1 0 0 0 0
47 50 1 0 0 0 0
46 51 1 0 0 0 0
1 52 1 0 0 0 0
52 34 1 0 0 0 0
53 54 1 0 0 0 0
25 53 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 53 54
M SBL 1 1 59
M SMT 1 CH2OH
M SBV 1 59 -0.7571 -0.1674
S SKP 5
ID FL5FACGL0048
FORMULA C33H40O21
EXACTMASS 772.206208342
AVERAGEMASS 772.6581
SMILES C(O6)(C(O)C(C(O)C6CO)O)OC(=C(c(c5)ccc(c5O)O)4)C(c(c(O)3)c(O4)cc(c3)OC(C1O)OC(COC(C2O)OC(C)C(C(O)2)O)C(C1O)O)=O
M END