Mol:FL5FAAGL0048
From Metabolomics.JP
(Difference between revisions)
| Line 1: | Line 1: | ||
| − | + | ||
| − | + | ||
| − | Copyright: ARM project http://www.metabolome.jp/ | + | Copyright: ARM project http://www.metabolome.jp/ |
| − | 52 57 0 0 0 0 0 0 0 0999 V2000 | + | 52 57 0 0 0 0 0 0 0 0999 V2000 |
| − | -4.2065 1.1898 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.2065 1.1898 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.2065 0.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.2065 0.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.5055 -0.0243 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.5055 -0.0243 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.8045 0.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.8045 0.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.8045 1.1898 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.8045 1.1898 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.5055 1.5944 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.5055 1.5944 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1036 -0.0243 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1036 -0.0243 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4026 0.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4026 0.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -1.4026 1.1898 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -1.4026 1.1898 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1036 1.5944 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1036 1.5944 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -2.1036 -0.6554 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -2.1036 -0.6554 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6851 1.6059 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6851 1.6059 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0294 1.1934 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0294 1.1934 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7438 1.6059 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7438 1.6059 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.7438 2.4309 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.7438 2.4309 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0294 2.8433 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0294 2.8433 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6851 2.4309 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6851 2.4309 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -3.5055 -0.6334 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -3.5055 -0.6334 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.5084 2.8723 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.5084 2.8723 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -4.8302 1.5944 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -4.8302 1.5944 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6928 -0.1356 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6928 -0.1356 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1679 -1.2926 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1679 -1.2926 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.7758 -1.0398 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.7758 -1.0398 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.2734 -1.8776 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.2734 -1.8776 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.2881 -2.8599 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.2881 -2.8599 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6556 -3.1128 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6556 -3.1128 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.1532 -2.2749 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.1532 -2.2749 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.2656 -0.5073 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.2656 -0.5073 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.8942 -2.6062 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.8942 -2.6062 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.6378 -3.6444 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.6378 -3.6444 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.2687 0.1242 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.2687 0.1242 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.9068 -0.3856 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.9068 -0.3856 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.9174 -0.3856 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.9174 -0.3856 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.4743 0.1919 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.4743 0.1919 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.3812 0.5838 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.3812 0.5838 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.9068 0.1506 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.9068 0.1506 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.9174 0.1134 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.9174 0.1134 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.2082 3.3568 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.2082 3.3568 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.7658 2.6210 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.7658 2.6210 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.5684 2.9331 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.5684 2.9331 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.3430 2.9415 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.3430 2.9415 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.7801 3.5045 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.7801 3.5045 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 3.0779 3.1339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 3.0779 3.1339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.9187 2.5773 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.9187 2.5773 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.1947 2.4690 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.1947 2.4690 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.8302 3.0623 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.8302 3.0623 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 0.0956 -4.0045 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 0.0956 -4.0045 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | -0.6395 -4.7393 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | -0.6395 -4.7393 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.5702 3.5015 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.5702 3.5015 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 4.3340 4.7393 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 4.3340 4.7393 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1.9174 -1.0459 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 | + | 1.9174 -1.0459 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 2.8177 -1.5657 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 | + | 2.8177 -1.5657 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0 |
| − | 1 2 1 0 0 0 0 | + | 1 2 1 0 0 0 0 |
| − | 2 3 2 0 0 0 0 | + | 2 3 2 0 0 0 0 |
| − | 4 5 2 0 0 0 0 | + | 4 5 2 0 0 0 0 |
| − | 5 6 1 0 0 0 0 | + | 5 6 1 0 0 0 0 |
| − | 6 1 2 0 0 0 0 | + | 6 1 2 0 0 0 0 |
| − | 4 7 1 0 0 0 0 | + | 4 7 1 0 0 0 0 |
| − | 7 8 1 0 0 0 0 | + | 7 8 1 0 0 0 0 |
| − | 8 9 2 0 0 0 0 | + | 8 9 2 0 0 0 0 |
| − | 9 10 1 0 0 0 0 | + | 9 10 1 0 0 0 0 |
| − | 10 5 1 0 0 0 0 | + | 10 5 1 0 0 0 0 |
| − | 7 11 2 0 0 0 0 | + | 7 11 2 0 0 0 0 |
| − | 9 12 1 0 0 0 0 | + | 9 12 1 0 0 0 0 |
| − | 12 13 2 0 0 0 0 | + | 12 13 2 0 0 0 0 |
| − | 13 14 1 0 0 0 0 | + | 13 14 1 0 0 0 0 |
| − | 14 15 2 0 0 0 0 | + | 14 15 2 0 0 0 0 |
| − | 15 16 1 0 0 0 0 | + | 15 16 1 0 0 0 0 |
| − | 16 17 2 0 0 0 0 | + | 16 17 2 0 0 0 0 |
| − | 17 12 1 0 0 0 0 | + | 17 12 1 0 0 0 0 |
| − | 3 18 1 0 0 0 0 | + | 3 18 1 0 0 0 0 |
| − | 19 15 1 0 0 0 0 | + | 19 15 1 0 0 0 0 |
| − | 20 1 1 0 0 0 0 | + | 20 1 1 0 0 0 0 |
| − | 21 8 1 0 0 0 0 | + | 21 8 1 0 0 0 0 |
| − | 4 3 1 0 0 0 0 | + | 4 3 1 0 0 0 0 |
| − | 22 23 1 0 0 0 0 | + | 22 23 1 0 0 0 0 |
| − | 23 24 1 1 0 0 0 | + | 23 24 1 1 0 0 0 |
| − | 24 25 1 1 0 0 0 | + | 24 25 1 1 0 0 0 |
| − | 26 25 1 1 0 0 0 | + | 26 25 1 1 0 0 0 |
| − | 26 27 1 0 0 0 0 | + | 26 27 1 0 0 0 0 |
| − | 27 22 1 0 0 0 0 | + | 27 22 1 0 0 0 0 |
| − | 22 28 1 0 0 0 0 | + | 22 28 1 0 0 0 0 |
| − | 27 29 1 0 0 0 0 | + | 27 29 1 0 0 0 0 |
| − | 26 30 1 0 0 0 0 | + | 26 30 1 0 0 0 0 |
| − | 23 21 1 0 0 0 0 | + | 23 21 1 0 0 0 0 |
| − | 31 32 1 1 0 0 0 | + | 31 32 1 1 0 0 0 |
| − | 32 33 1 1 0 0 0 | + | 32 33 1 1 0 0 0 |
| − | 34 33 1 1 0 0 0 | + | 34 33 1 1 0 0 0 |
| − | 34 35 1 0 0 0 0 | + | 34 35 1 0 0 0 0 |
| − | 35 31 1 0 0 0 0 | + | 35 31 1 0 0 0 0 |
| − | 32 36 1 0 0 0 0 | + | 32 36 1 0 0 0 0 |
| − | 33 37 1 0 0 0 0 | + | 33 37 1 0 0 0 0 |
| − | 31 28 1 0 0 0 0 | + | 31 28 1 0 0 0 0 |
| − | 38 39 1 1 0 0 0 | + | 38 39 1 1 0 0 0 |
| − | 39 40 1 1 0 0 0 | + | 39 40 1 1 0 0 0 |
| − | 41 40 1 1 0 0 0 | + | 41 40 1 1 0 0 0 |
| − | 41 42 1 0 0 0 0 | + | 41 42 1 0 0 0 0 |
| − | 42 43 1 0 0 0 0 | + | 42 43 1 0 0 0 0 |
| − | 43 38 1 0 0 0 0 | + | 43 38 1 0 0 0 0 |
| − | 39 44 1 0 0 0 0 | + | 39 44 1 0 0 0 0 |
| − | 40 45 1 0 0 0 0 | + | 40 45 1 0 0 0 0 |
| − | 19 38 1 0 0 0 0 | + | 19 38 1 0 0 0 0 |
| − | 41 46 1 0 0 0 0 | + | 41 46 1 0 0 0 0 |
| − | 47 48 1 0 0 0 0 | + | 47 48 1 0 0 0 0 |
| − | 25 47 1 0 0 0 0 | + | 25 47 1 0 0 0 0 |
| − | 49 50 1 0 0 0 0 | + | 49 50 1 0 0 0 0 |
| − | 42 49 1 0 0 0 0 | + | 42 49 1 0 0 0 0 |
| − | 51 52 1 0 0 0 0 | + | 51 52 1 0 0 0 0 |
| − | 33 51 1 0 0 0 0 | + | 33 51 1 0 0 0 0 |
| − | M STY 1 1 SUP | + | M STY 1 1 SUP |
| − | M SLB 1 1 1 | + | M SLB 1 1 1 |
| − | M SAL 1 2 47 48 | + | M SAL 1 2 47 48 |
| − | M SBL 1 1 53 | + | M SBL 1 1 53 |
| − | M SMT 1 CH2OH | + | M SMT 1 CH2OH |
| − | M SBV 1 53 -0.3837 1.1446 | + | M SBV 1 53 -0.3837 1.1446 |
| − | M STY 1 2 SUP | + | M STY 1 2 SUP |
| − | M SLB 1 2 2 | + | M SLB 1 2 2 |
| − | M SAL 2 2 49 50 | + | M SAL 2 2 49 50 |
| − | M SBL 2 1 55 | + | M SBL 2 1 55 |
| − | M SMT 2 CH2OH | + | M SMT 2 CH2OH |
| − | M SBV 2 55 -0.7901 0.0029 | + | M SBV 2 55 -0.7901 0.0029 |
| − | M STY 1 3 SUP | + | M STY 1 3 SUP |
| − | M SLB 1 3 3 | + | M SLB 1 3 3 |
| − | M SAL 3 2 51 52 | + | M SAL 3 2 51 52 |
| − | M SBL 3 1 57 | + | M SBL 3 1 57 |
| − | M SMT 3 CH2OH | + | M SMT 3 CH2OH |
| − | M SBV 3 57 0.0000 0.6603 | + | M SBV 3 57 0.0000 0.6603 |
| − | S SKP 5 | + | S SKP 5 |
| − | ID FL5FAAGL0048 | + | ID FL5FAAGL0048 |
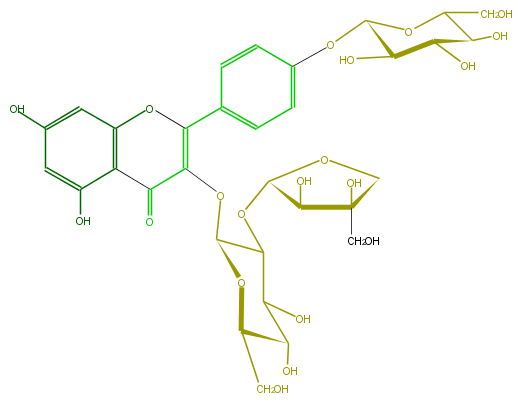
| − | FORMULA C32H38O20 | + | FORMULA C32H38O20 |
| − | EXACTMASS 742.1956436559999 | + | EXACTMASS 742.1956436559999 |
| − | AVERAGEMASS 742.63212 | + | AVERAGEMASS 742.63212 |
| − | SMILES O(C(C4=O)=C(c(c5)ccc(OC(O6)C(C(C(O)C6CO)O)O)c5)Oc(c34)cc(cc3O)O)C(C(OC(O2)C(O)C(O)(CO)C2)1)OC(C(O)C(O)1)CO | + | SMILES O(C(C4=O)=C(c(c5)ccc(OC(O6)C(C(C(O)C6CO)O)O)c5)Oc(c34)cc(cc3O)O)C(C(OC(O2)C(O)C(O)(CO)C2)1)OC(C(O)C(O)1)CO |
M END | M END | ||
| − | |||
Latest revision as of 09:00, 14 March 2009
Copyright: ARM project http://www.metabolome.jp/
52 57 0 0 0 0 0 0 0 0999 V2000
-4.2065 1.1898 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-4.2065 0.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.5055 -0.0243 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.8045 0.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.8045 1.1898 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.5055 1.5944 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.1036 -0.0243 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.4026 0.3804 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-1.4026 1.1898 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-2.1036 1.5944 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-2.1036 -0.6554 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.6851 1.6059 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0294 1.1934 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7438 1.6059 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.7438 2.4309 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.0294 2.8433 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.6851 2.4309 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-3.5055 -0.6334 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.5084 2.8723 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-4.8302 1.5944 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.6928 -0.1356 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.1679 -1.2926 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.7758 -1.0398 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.2734 -1.8776 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
-0.2881 -2.8599 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.6556 -3.1128 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.1532 -2.2749 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.2656 -0.5073 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.8942 -2.6062 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.6378 -3.6444 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.2687 0.1242 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
0.9068 -0.3856 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.9174 -0.3856 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.4743 0.1919 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
1.3812 0.5838 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.9068 0.1506 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.9174 0.1134 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
2.2082 3.3568 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.7658 2.6210 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.5684 2.9331 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3430 2.9415 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.7801 3.5045 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
3.0779 3.1339 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.9187 2.5773 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.1947 2.4690 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.8302 3.0623 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
0.0956 -4.0045 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
-0.6395 -4.7393 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
4.5702 3.5015 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
4.3340 4.7393 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1.9174 -1.0459 0.0000 C 0 0 0 0 0 0 0 0 0 0 0 0
2.8177 -1.5657 0.0000 O 0 0 0 0 0 0 0 0 0 0 0 0
1 2 1 0 0 0 0
2 3 2 0 0 0 0
4 5 2 0 0 0 0
5 6 1 0 0 0 0
6 1 2 0 0 0 0
4 7 1 0 0 0 0
7 8 1 0 0 0 0
8 9 2 0 0 0 0
9 10 1 0 0 0 0
10 5 1 0 0 0 0
7 11 2 0 0 0 0
9 12 1 0 0 0 0
12 13 2 0 0 0 0
13 14 1 0 0 0 0
14 15 2 0 0 0 0
15 16 1 0 0 0 0
16 17 2 0 0 0 0
17 12 1 0 0 0 0
3 18 1 0 0 0 0
19 15 1 0 0 0 0
20 1 1 0 0 0 0
21 8 1 0 0 0 0
4 3 1 0 0 0 0
22 23 1 0 0 0 0
23 24 1 1 0 0 0
24 25 1 1 0 0 0
26 25 1 1 0 0 0
26 27 1 0 0 0 0
27 22 1 0 0 0 0
22 28 1 0 0 0 0
27 29 1 0 0 0 0
26 30 1 0 0 0 0
23 21 1 0 0 0 0
31 32 1 1 0 0 0
32 33 1 1 0 0 0
34 33 1 1 0 0 0
34 35 1 0 0 0 0
35 31 1 0 0 0 0
32 36 1 0 0 0 0
33 37 1 0 0 0 0
31 28 1 0 0 0 0
38 39 1 1 0 0 0
39 40 1 1 0 0 0
41 40 1 1 0 0 0
41 42 1 0 0 0 0
42 43 1 0 0 0 0
43 38 1 0 0 0 0
39 44 1 0 0 0 0
40 45 1 0 0 0 0
19 38 1 0 0 0 0
41 46 1 0 0 0 0
47 48 1 0 0 0 0
25 47 1 0 0 0 0
49 50 1 0 0 0 0
42 49 1 0 0 0 0
51 52 1 0 0 0 0
33 51 1 0 0 0 0
M STY 1 1 SUP
M SLB 1 1 1
M SAL 1 2 47 48
M SBL 1 1 53
M SMT 1 CH2OH
M SBV 1 53 -0.3837 1.1446
M STY 1 2 SUP
M SLB 1 2 2
M SAL 2 2 49 50
M SBL 2 1 55
M SMT 2 CH2OH
M SBV 2 55 -0.7901 0.0029
M STY 1 3 SUP
M SLB 1 3 3
M SAL 3 2 51 52
M SBL 3 1 57
M SMT 3 CH2OH
M SBV 3 57 0.0000 0.6603
S SKP 5
ID FL5FAAGL0048
FORMULA C32H38O20
EXACTMASS 742.1956436559999
AVERAGEMASS 742.63212
SMILES O(C(C4=O)=C(c(c5)ccc(OC(O6)C(C(C(O)C6CO)O)O)c5)Oc(c34)cc(cc3O)O)C(C(OC(O2)C(O)C(O)(CO)C2)1)OC(C(O)C(O)1)CO
M END